Autophagy is a natural mechanism in which the cell removes and degrades cellular components with autolysosomes. It is a popular research area because autophagy is related to many physical and pathological processes. The 2016 Nobel Prize in Physiology is granted to Yoshinori Ohsumi for his contribution in autophagy. In autophagy studies, LC3-I and LC3-II detection is a must-have experiment to track the autophagy process. Therefore, we would like to share five important notes for quantifying autophagy with LC3.
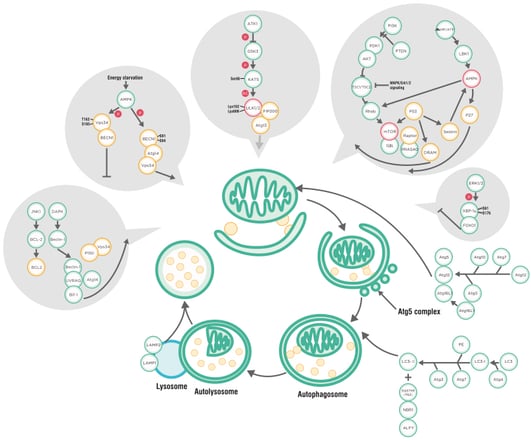
LC3 is a key protein in the autophagy pathway. Once synthesized, cytoplasmic LC3 is processed by Atg4 at the C-terminal, forming cytoplasmic LC3-I (16 KDa). Upon autophagy signal, LC3-I is conjugated by ubiquitin-like proteins Atg7 and Atg3 to the lipid phosphatidylethanolamine (PE), generating lipidated LC3-II (14 KDa). The lipidated LC3-II binds to both inner and outer membranes of autophagosomes, with the former being degraded after fusion with lysosomes; whereas LC3 on the outer membrane is deconjugated by ATG4 and returns to the cytosol. Thus, LC3-I to LC3-II conversion and lysosomal degradation of LC3-II reflect the progression of autophagy, and immunoblotting can easily be used to monitor changes in LC3 amount.
Autophagy Pathway
Click here to download full size poster
1. Choose the correct LC3 isoform(s) for your research
Mammalian LC3 family has four isoforms, LC3A, LC3B, LC3B2 and LC3C. The LC3C is believed to be poorly expressed or not expressed in most normal tissues; while both LC3A and LC3B are differentially expressed in most normal tissues and cells. The choice of LC3 isoform(s) should be based on sample species as well as types of tissues and cells. The following two figures shows the mRNA expression difference of LC3A and LC3B in mice. (Source: Biogps)
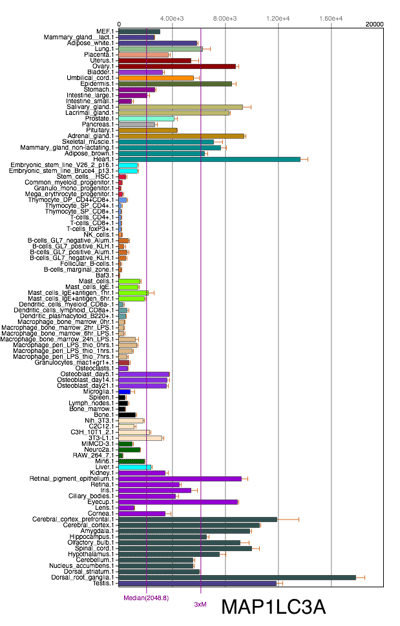
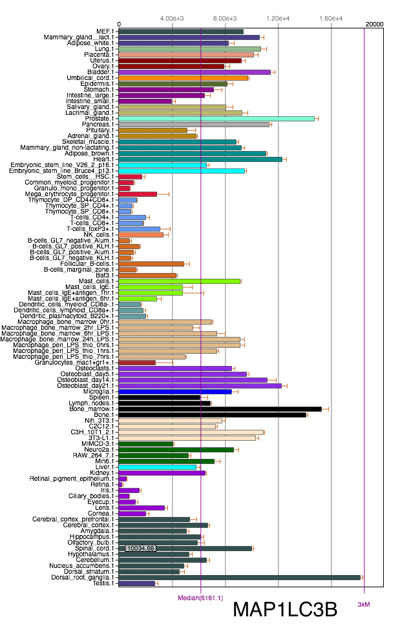


Click image to view original
2. Prevent lysosomal degradation of LC3-II
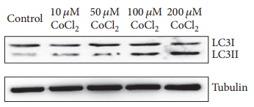
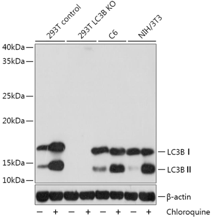
As described in the autophagy process above, increased LC3-II could reflect either increased autophagosome formation or decreased autophagosome degradation through fusion with lysosomes, which means the level of LC3-II at a specific time point is insufficient for an overall estimation of autophagic flux. Thus, we suggest to detect LC3-II turnover by WB in both presence and absence of lysosomal degradation. Common lysosomal degradation inhibitors include chloroquine and bafilomycin A1. Chloroquine inhibits lysosome functions via increase lysosomal PH; bafilomycin A1 is specifically targeting V-ATPase and thus inhibits autophagosome-lysosome fusion.
3. Use housekeeping proteins
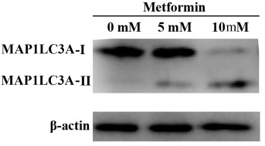
Currently, epitope sites of most LC3 antibodies are chosen at the N-terminal. However, the PE unit conjugation might induce protein conformational change at the N-terminal, and consequently increase the sensitivity of LC3 antibodies. Thus, the intense color band of LC3-II does not necessarily indicate high autophagy level of the sample. Also, LC3-I is particularly unstable to freeze-thaw cycles and likely to degrade in SDS loading buffer. Therefore, instead of LC3-II to LC3-II conversion ratio, measuring the levels of LC3-II to appropriate housekeeping proteins (e.g., β-actin) is likely to be a more reliable method.
4. Detach LC3 from membranes
LC3 is a protein that can be lipidated and often localized on membranes. To facilitate its detachment from membranes, you can sonicate the sample on ice to lyse the cells completely. Sonicate for a total of three times using 35%-40% of the max power (max power of different equipments can be adjusted to around 200 W), with 10 seconds for each sonication and 10 seconds between sonications.
5. Apply high-percentage resolving gels
It is of note that although the actual molecular weight of LC3-II (PE-conjugated) is larger than that of LC3-I (16 KDa), LC3-II migrates faster than LC3-I on SDS-PAGE due to its extreme hydrophobicity, reaching approximately 14KDa. Due to the small molecular weight difference (2 KDa) between LC3-II and LC3-I, high-percentage gels (12% to 15%) are desired to successfully separate them. For wet transfer, use 0.22 μm pore size membrane filter and transfer at the constant voltage of 90 V for 70 min. Apply 2.5A for 7 min when using a semi-dry blotter.
ABclonal Featured LC3 Antibodies
LC3B Rabbit mAb (A19665)  |
LC3B Rabbit pAb (A7198)  |
MAP1LC3A Rabbit pAb (A11438)  |
LC3A/LC3B Rabbit pAb (A5618)  |
 |
Publication
|
 |
References
Barth, Sandra, Danielle Glick, and Kay F Macleod. "Autophagy: Assays and Artifacts." The Journal of pathology 221.2 (2010): 117-24. Print.
Chan, Leo Li-Ying, et al. "A Novel Image-Based Cytometry Method for Autophagy Detection in Living Cells." Autophagy 8.9 (2012): 1371-82. Print.
Singh, Bindu, and Sangeeta Bhaskar. "Methods for Detection of Autophagy in Mammalian Cells." Stem Cells and Aging. Springer, 2018. 245-58. Print.
Zhang, Ziyan, Rajat Singh, and Michael Aschner. "Methods for the Detection of Autophagy in Mammalian Cells." Current protocols in toxicology 69.1 (2016): 20.12. 1-20.12. 26. Print.