1. Determine the mRNA capture type you need

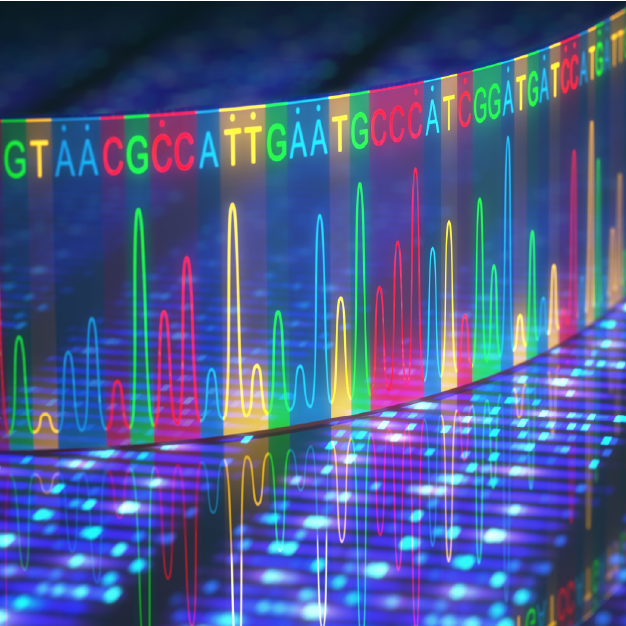
PolyA RNA capture targets total RNA samples from eukaryotes such as animals, plants, and fungi, because mature mRNA in these types of samples has a polyA tail structure at the 3' end. Hence, magnetic beads chelated with oligo dT are used to capture polyA RNA through annealing, in a quick and easy manner.
PolyA RNA capture modules can be used to quantify gene expression, alternative splicing, fusion genes, single-nucleotide variant (SNV), and more.
rRNA depletion (H/M/R) module is mainly used for human, mouse, and rat total RNA. It removes cytoplasmic ribosomal RNA (28S, 18S, 5S, 5.8 S) and mitochondrial rRNA (12S, 16S) from the total RNA by probe hybridization and enzymatic hydrolysis. This method can enrich all RNA except rRNA and small RNA, and can analyze more long-chain non-coding RNA (lncRNA), circRNA and other non-polyA tail transcripts.
You can choose the appropriate mRNA capture method according to the experimental needs and ABclonal offers both mRNA capture modules.
2. RNA sample quality

After selecting the mRNA capturing method, purification of RNA samples is paramount to building a RNA library.
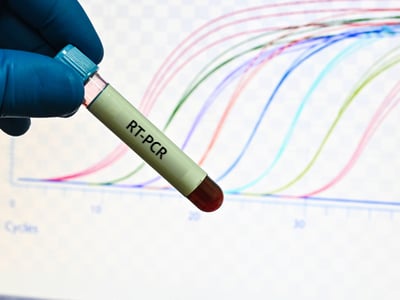
RNA quality assessment is the first step in RNA-seq construction. The usual method for assessing RNA quality is to use Agilent's Bioanalyzer, which gives an RNA integrity score based on the degree of degradation of the RNA sample, expressed as RIN. The RIN value ranges from 0 to 10. A higher value indicates better RNA integrity, and conversely, the more severe the sample is degraded.
ABclonal mRNA-seq lib prep requires RNA to have a RIN value of 7 or higher. If the sample is degraded, it is easy to cause 3' end preference, and it is difficult to comprehensively analyze the SNV and structure of the gene.
The ABclonal Whole RNA-seq lib prep kit is not only suitable for intact RNA samples, but also for degraded RNA samples such as FFPE RNA samples.
3. Strand-specific libraries vs. non-strand specificity

After the RNA sample is detected, there is the choice of constructing a chain-specific or non-chain-specific library.
The transcription of genes can be derived from the positive or negative strands of DNA. Chain-specific libraries and common transcriptome libraries differ in that chain-specific libraries enable you to differentiate between positive and negative strands. By doing so, the number of transcripts and lncRNA can both be quantified/analyzed more accurately.
ABclonal has two types of mRNA library preparation kits, mRNA-seq Lib Prep Kit and Stranded mRNA-seq Lib Prep Kit. The strand-specific library construction kit uses dUTP instead of dTTP to label second-strand cDNA during synthesis. Uracil-DNA glycosylase (UDG) is used to hydrolyze the strand before PCR enrichment to ensure that the final sequencing data is from first strand cDNA. In addition, actinomycin D is added to the first strand cDNA synthesis to inhibit the binding of reverse transcriptase to the DNA template and to avoid the synthesis of a pseudo-negative strand – enhancing the strand specificity of the sequencing data.
4. Truncated adapter vs. full-length adapter

The truncated adapter is based on the full length 'Y'-shaped adapters from Illumina, which cuts off the non-complementary split-section area, leaving a part of the complementary area and a small part of the split area. It is found that truncated adapters better connect to 3A dsDNA and are less likely to form a linker dimer compared to full-length adapters. The truncated portion is complemented in the library enrichment step by the tailed primer. This ensures that the constructed library can carry the complete P5 and P7 sequences, ensuring efficient hybridization with the sequencing chip and completing the sequencing process.
Apart from optimizing the DNA library preparation buffers and enzymes to enhance library preparation processes, ABclonal also uses truncated adapters to improve efficiency.
5. PCR mix
The choice of PCR polymerase also plays a crucial role in RNA library construction using next-generation sequencing. High amplification efficiency allows you to obtain an efficient library yield with the minimum number of cycles, thereby reducing the generation of duplication in sequencing data. A highly specific PCR polymerase ensures the library specificity and reduces the production of non-specific products. A high sensitivity ensures that low-copy genes can be amplified well. The high fidelity 2 × PCR mix included in ABclonal’s RNA-seq library is not only easy to use, but also has high amplification efficiency, high PCR specificity and high sensitivity. It is the best choice for RNA-seq library construction.