 A common experimental strategy in studying the effects of a specific protein in cells or organisms is to remove it. One can determine the physiological outcomes in the absence of that protein to ascertain its relative importance in maintaining normal functions, or in some cases, to note that it is dispensable or redundant and might have a backup within the cell to take up the slack. Some targeted techniques include RNA interference (RNAi) and CRISPR-based gene editing, and in many cases, it is possible to generate knockout cell lines or even organisms, like mice, that cannot express a specific protein. But when those strategies are not feasible for the experiment at hand, what is one to do?
A common experimental strategy in studying the effects of a specific protein in cells or organisms is to remove it. One can determine the physiological outcomes in the absence of that protein to ascertain its relative importance in maintaining normal functions, or in some cases, to note that it is dispensable or redundant and might have a backup within the cell to take up the slack. Some targeted techniques include RNA interference (RNAi) and CRISPR-based gene editing, and in many cases, it is possible to generate knockout cell lines or even organisms, like mice, that cannot express a specific protein. But when those strategies are not feasible for the experiment at hand, what is one to do?
Direct Protein Degradation
DNA knockouts, RNAi, and CRISPR-Cas9 all act at the genome or transcriptional level, but that acts indirectly to prevent the production of the target protein. Therefore, it takes some time for RNAi experiments with small-interfering RNA (siRNA) to incubate, to give the cell time to eliminate the target mRNA and thus prevent the target protein from being expressed, as well as for the cell to clear out the existing inventory of said protein.
 This prolonged mechanism motivated the discovery of a novel technique that would directly target and degrade the desired protein immediately. In the original resource article that Gavin showed me, the authors presented a technique termed “Trim-Away” that uses ubiquitination and proteasome-mediated degradation in concert with a specific antibody to ensure precise degradation of the target protein. 1
This prolonged mechanism motivated the discovery of a novel technique that would directly target and degrade the desired protein immediately. In the original resource article that Gavin showed me, the authors presented a technique termed “Trim-Away” that uses ubiquitination and proteasome-mediated degradation in concert with a specific antibody to ensure precise degradation of the target protein. 1
The researchers were also previously aware of a cytosolic protein called TRIM21 that acts as an E3 ubiquitin ligase, which promotes the degradation of ubiquitinated proteins by the proteasome. TRIM21 appears to also bind tightly to the Fc domain of antibodies and is expressed in a wide range of cell types to degrade pathogens like viruses and bacteria. 1
The Trim-Away Mechanism
 In developing Trim-Away, the researchers wanted to employ antibodies due to their robustness and ability to recognize and strongly bind specific amino acid sequences, which would facilitate the targeting of post-translational modifications and point mutations. The nigh ubiquitous expression of TRIM21 along with its apparent Fc-binding ability allowed the conception of the new technique to target and destroy a protein of interest in rapid order. (Figure 1)
In developing Trim-Away, the researchers wanted to employ antibodies due to their robustness and ability to recognize and strongly bind specific amino acid sequences, which would facilitate the targeting of post-translational modifications and point mutations. The nigh ubiquitous expression of TRIM21 along with its apparent Fc-binding ability allowed the conception of the new technique to target and destroy a protein of interest in rapid order. (Figure 1)
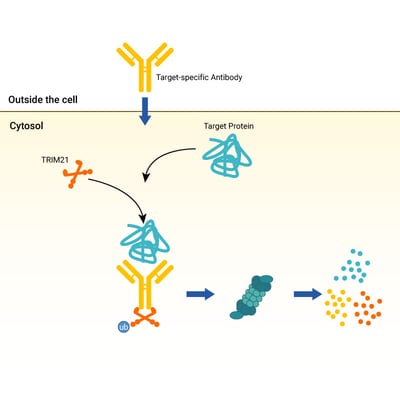
The procedure is performed stepwise with the introduction of exogenous TRIM21 (though the endogenous level of TRIM21 seems sufficient), the subsequent addition of an antibody specific to the protein of interest, and finally, the coordination of TRIM21 with the antibody-protein complex to drive its degradation. 1 (Figure 1) When initially testing their technique, the researchers noted that Trim-Away works with a variety of different proteins localized to many different regions of the cell, whether they are floating around as individual units or incorporated into larger complexes or structures. The duration of degradation also coincided with the amount of antibody used.
Figure 1 (above, right): The Trim-Away mechanism requires either the endogenous TRIM21 protein or for the user to transfect in exogenous TRIM21. A target specific antibody will bind the desired protein, which then complexes with TRIM21 to be directed to the proteasome for degradation.
Applications of Trim-Away
Because antibodies are huge (usually around 150 kDa in size), the worry is that they would not be able to penetrate the cell membrane (although electroporation in the original experiment seems to work well),1 tissues, or might have problems binding to hidden or sterically hindered domains. A group in China proposed an alternative approach that keeps the basic Trim-Away mechanism in place but uses a smaller antibody-based structure called a nanobody. In this methodology, the nanobody, which is a recombinant heavy-chain-only epitope-binding structure, is fused to a truncated version of TRIM21 that maintains its ubiquitination activity, which the authors called a TRIMbody. 2 With a smaller nanobody (about 15 kDa) and a shorter version of the Trim21 protein, the fusion is less than half as large as a full-sized antibody and was postulated to be more amenable to hard-to-reach epitopes in target proteins. The authors only tested this with a customized EGFP construct, so it would be interesting to see how this works with TRIMbody fusions against a wider range of natural proteins.
Due to the rapidity of degradation via the Trim-Away technique, the dream of disease researchers is to use this new technology as another delivery vehicle for targeted therapies. In Huntington disease, for example, the primary cause is the aggregation of a mutated huntingin (mHTT) protein in nerve cells. Shortly after Trim-Away was reported, mHTT was tested to see if the technique was viable. While Trim-Away could generate protein knockdown cell lines for functional research, it has yet to be used in actual therapies. 3 Part of the challenge, like other methodologies involving genetic approaches like RNAi and CRISPR, is the selective delivery to the affected regions, especially with the need to deliver either a large antibody or even a TRIMbody.
How ABclonal Can Help
![]() As you may know, ABclonal boasts a catalog comprising thousands of high-quality antibody products, many of which should be compatible with the "conventional" Trim-Away technique. If you wish to give the modified TRIMbody technique a go, contact us for our customized antibody expression services in which we might be able to provide you with constructs that you can then incorporate into your own unique TRIMbody. Our molecular biology products will also be useful to you as you work toward making Trim-Away and TRIMbody techniques more amenable to targeted therapies in addition to studies in cell lines.
As you may know, ABclonal boasts a catalog comprising thousands of high-quality antibody products, many of which should be compatible with the "conventional" Trim-Away technique. If you wish to give the modified TRIMbody technique a go, contact us for our customized antibody expression services in which we might be able to provide you with constructs that you can then incorporate into your own unique TRIMbody. Our molecular biology products will also be useful to you as you work toward making Trim-Away and TRIMbody techniques more amenable to targeted therapies in addition to studies in cell lines.
References
- Clift et al. (2017) “A Method for the Acute and Rapid Degradation of Endogenous Proteins.” Cell 171(7):1692-1706.
- Chen et al. (2021) “A Promising Intracellular Protein-Degradation Strategy: TRIMbody-Away Technique Based on Nanobody Fragment.” Biomolecules 11(10):1512 (Epub).
- Jarosinska OD and Rudiger SGD (2021) “Molecular Strategies to Target Protein Aggregation in Huntington’s disease.” Front Mol Biosci 8:769184 (Epub).