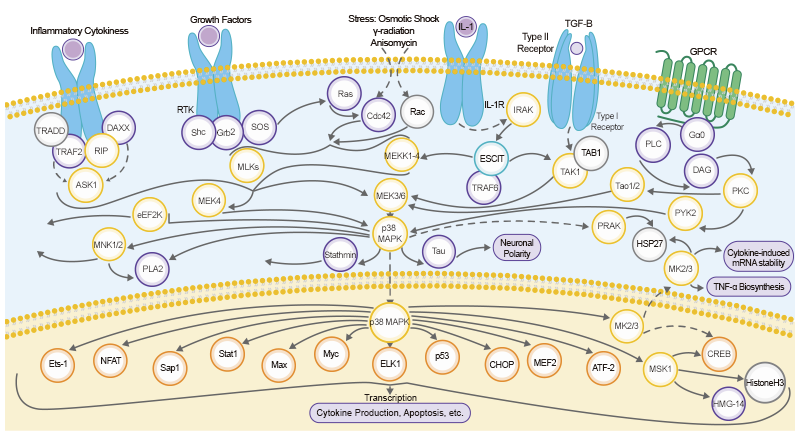
Proteins known as transcription factors play a crucial role in gene regulation by activating, enhancing, and even silencing a gene’s expression. Many textbooks and resources compare transcription factors (TFs) to something like an on/off switch for gene transcription. However, it is a bit more complicated than just turning gene expression on or off. Various properties (e.g. binding affinity, specificity, and genetic variance of binding sites) impact the binding of TFs to DNA, thereby altering gene expression. To study transcription and how it is regulated, scientists study TF-DNA interactions on a genome-wide level.
Various techniques, both in vivo and in vitro have been developed for scientists to characterize TF-DNA binding. However, no technique is yet perfect and researchers are still working to improve these methods. One major shortcoming in existing methods is that they do not account for the impacts of cellular environments. As a result, characterizing TF-DNA interactions remains a challenge-- only a handful of TFs have been characterized well enough to predict DNA binding.
Existing Methods
Numerous experimental methods, both in vivo and in vitro have been developed to characterize TF-DNA interactions, but they each have their own shortcomings.
In vivo
In vivo-based methods identify TF binding sites and the biological context of DNA-specific interactions. When it comes to binding site resolution however, in vivo methods have lower resolution than their in vitro counterparts. The resolution is typically between 100 and 500 bp. In vivo methods are also incapable of distinguishing between direct or indirect interactions.
The most commonly used in vivo method is called chromatin immunoprecipitation (ChIP) which is used to study genome-wide TF binding. It works by cross-linking transcription factors to DNA. The complexes are fixed, sheared, and immunoprecipitated with a TF-specific antibody. Modified ChIP methodologies also exist such as ChIP-chip and ChIP-seq.
In vitro
In vitro methods identify TF binding sites, binding energy landscapes, and the biophysical parameters governing these binding events. The ability to distinguish direct and indirect interactions is a major advantage of in vitro methods. Higher resolution, often to the nucleotide level, is also feasible with the majority of these methods. However, as this articles mentions above, in vitro methods do not account for cell-specific impacts since they use purified or in vitro produced protein samples. In vitro methods include:
- Systematic evolution of ligands by exponential enrichment (SELEX)
- DNA immunoprecipitation (DIP- chip)
- Protein-binding microarrays (PBM)
New Method: NextPBM
To improve on current techniques, a group of scientists at Boston University recently developed nextPBM, a method that accounts for cell-specific impacts such as post-translational modifications and cooperative cofactors. Demonstrated with ABclonal’s FLI1 antibody, the method modifies the existing in vitro technique, PBM. The technique incorporates a computational framework, which can discover cell-specific cofactors, screen for synthetic cooperative DNA elements, and characterize TF cooperativity.
If you're looking for other helpful reads regarding proteins, be sure to check out our other related blogs here.
.jpg)