Jan 23, 2025 12:42:04 PM by Team - ABclonal
ABclonal x YCharOS: Setting the Gold Standard in Antibody Characterization
Detailed Guideline on Pyroptosis Research
It is evident that the gasdermin family is central to pyroptosis. This is because the gasdermin family members are the executioners of pyroptosis, and their pore-forming function is a necessary condition for pyroptosis to occur. This family generally includes a cytotoxic N-terminal domain and a C-terminal inhibitory domain. After cleavage, the N-terminal domain is released and can assemble into pores in the membrane. Gasdermin pores disrupt the integrity of the cell membrane, leading to inflammatory cell death, with cell contents, including inflammatory cytokines, being released into the extracellular space.
In the 1980s and 1990s, scientists discovered that macrophages exposed to bacterial toxins or infections underwent a unique form of cell death that required the activation of caspase-1. For years, this process was mistakenly classified as apoptosis, a non-inflammatory and orderly form of cell death. However, unlike apoptosis, this newly observed cell death was marked by swelling, rupture, and the release of inflammatory signals, indicating a far more chaotic and immune-activating process. It wasn’t until 2001 that Cookson BT and Brennan MA officially named this inflammatory form of cell death “pyroptosis”. [2] Since then, research has revealed that pyroptosis plays a key role in immune defense by promoting inflammation in response to infections. The discovery of gasdermin D (GSDMD), a protein that forms pores in the cell membrane, further clarified the mechanism of pyroptosis. Today, it’s understood that pyroptosis occurs in various cell types and is critical in both fighting infections and contributing to inflammatory diseases when dysregulated.
Although the phenomenon of cell death had been known for centuries, even briefly described in the 19th century, the explosion of research and technology in the latter half of the 20th century led to greater and more nuanced discoveries that have provided insights into many physiological processes including tissue development, maintenance, metabolism, and disease states including cancer. The many accomplishments in programmed cell death research have improved our understanding and development of targeted therapeutics, and some of these milestones were recognized by Nobel Prizes for apoptosis and autophagy. As scientists continue to elucidate new cell death pathways and their interplay with other pathways, let's take a look at what we know so far and what new findings have come out.
Every August, we observe National Immunization Awareness Month (NIAM), an event that educates and encourages everyone to keep up with their vaccinations. While healthcare providers continue to play important roles in education and supporting public health, the rest of us can read up on what vaccines are needed, vaccination schedules, and provide outreach to ensure that we and our neighbors remain safe and healthy from preventable diseases.
Aug 5, 2024 2:35:29 PM by Team ABclonal
Targeting Epigenetic Marks: How Antibodies Empower Cancer Research
Introduction
In the realm of genetics, the discovery of DNA and its double helix structure led to the assumption that genetic sequences alone determine cell phenotypes. However, researchers began to observe cases where organisms with identical genetic information exhibited different traits. This realization gave birth to the field of epigenetics, which explores the reversible influence on gene expression without altering DNA sequences. Epigenetic mechanisms encompass DNA methylation, histone modification, chromatin remodeling, and the effects of noncoding RNA. These processes involve “writers,” “readers,” and “erasers” that add, recognize, or remove chemical groups from DNA or histones. The cooperation between the epigenome, transcription factors, noncoding RNAs, and external stimuli regulates gene expression in a temporary yet long-lasting manner. Understanding normal and abnormal epigenetic processes is crucial for comprehending diseases like cancer and developing potential treatments.
Having just moved a ludicrous amount of boxes and furniture into various U-Hauls and relocation tubes, I can feel all the literal weight of those decisions in my muscles and bones. Now that I'm back in Chicago, nursing my muscle soreness and the occasional bruise, I'm left thinking about the need for better muscular recovery and repair, which brings us to today's wonderful success story with an ABclonal customer as they added to our knowledge of myoblast differentiation and skeletal muscle development.
In the wake of new programs that produce artwork derived from existing media, ChatGPT, and even algorithms that can predict protein folding, it is evident that the age of artificial intelligence (AI) is upon us. In many cases, the AI programs and tools are far more advanced than we have previously seen, to the point where humanity can derive great benefit from AI while fearing how it may affect our society and livelihoods. While it is unlikely that we will be subjugated by our new robot overlords, it is still important to explore what has been done and remains possible through AI, and our considerations for its ethical usage.
Perhaps one of the best things about awareness months, even if they bring focus to maladies and situations that are not always pleasant, is to get people interested to learn about the topic, and if they're interested enough, they'll do something to help out. This is the case with amyotrophic lateral sclerosis, or ALS. Some time back, there was the Ice Bucket Challenge which went viral, involved numerous athletes and celebrities and common people, and raised a ton of money for research and to support the afflicted people around the world. Yet, although I know it is a motor neuron disease and that it has affected famous people like the late brilliant physicist Stephen Hawking and more recently, the respected baseball media personality and fellow UChicago alum Sarah Langs (she's not even that old!), I found that I knew remarkably little about the disease itself. So this May, which is ALS Awareness Month, let's learn a little bit more together.
Parkinson's Disease (PD) remains the second-most common neurodegenerative disorder behind Alzheimer's Disease, and the incidence of patients being diagnosed with PD will only rise as we all get older. With this in mind, much personnel and resources are dedicated to discovering more about this disease and to develop better treatments and management strategies to improve the livelihoods of those afflicted with PD. April is Parkinson's Awareness Month, and this is a perfect time for us not only to raise funds and awareness to help PD patients, but also to learn about how companies like ABclonal can help accelerate the research behind PD onset and progression. Here, we highlight studies using ABclonal products that were published within the past year that add new insights into Parkinson's research.
It is always a thrill and a privilege to share our customers' success stories with you, particularly when they feature ABclonal's products in their research publications. Over the past year, we have seen many citations of our reagents in multiple journals, showcasing the partnership ABclonal maintains as a trusted lab partner across a wide range of disciplines. In this entry, we will highlight some of the catalog and custom antibody products our valued customers used to generate recent publication-quality research that adds to our collective understanding of biology!
Jan 10, 2024 12:00:00 PM by Kin Leung
Diabetes, Obesity, and Accessibility: Considerations For GLP-1 Drugs
As the new year gets going, most people have already set some New Year's resolutions (here's some if you're of the science-y academic ilk) and chief among these is usually to exercise more and lose that holiday weight after the mounds of cookies and pie. It is perhaps not a coincidence that the latest Science Magazine Breakthrough of the Year recognizes GLP-1 drugs, which have been shown not only to be effective in managing diabetes, but also has significant impact on weight loss. The demand for these drugs has skyrocketed over the last few years as the word of their efficacy has spread, leading to supply chain issues that could adversely affect diabetes patients dependent on the medications. As the various GLP-1 drugs remain in short supply, it seems a good time to explore these drugs, their potential contributions to global human health, and what can be done to ensure they are accessible to those who need them most.
My wife put her studies on hold while I completed my PhD in cancer biology to take care of our son, but is now on the verge of getting her art history degree! One of her current classes is discussing the role of technology in art, and she stumbled across this older article that addresses the field of "bio-art," or art that represents a crossover between art and the biological sciences. I never really considered this as a contemporary art form, but I guess, separate from the illustrators or photographers who produce content for textbooks and science magazines, as well as the graphic designers who make the signal transduction diagrams and informational graphics for our references, bioscience has advanced to a point where we can manipulate cells in culture and living organisms to produce works of art that could be easily recognizable or more abstract. Since we are talking about living tissues being employed in the display of art, there are ethical considerations in play that we will explore in this article.
In 2022, Science Magazine awarded their Science Breakthrough of the Year to the NASA JW Space Telescope. In the time since, we have been mesmerized by the many new picture of our vast cosmos. Since ancient humans gazed up into the stars, we have always wondered what was out there in the universe, and we can find our fascination with the final frontier in our mythology, both from ages past and in contemporary science fiction. As humanity looks to return to the moon and establish further presence on Mars and beyond, let's take a look at our search for extraterrestrial life, if it is out there...or should I say, it's most likely out there!
Nov 15, 2023 11:54:38 AM by Kin Leung
The Magic Bullet: Current State of the Antibody-Drug Conjugate Market
The dream of biomedical researchers is to fine-tune their therapeutics to precisely target the specific illness or pathogen affecting their patient. Ever since Nobel laureate and oft-quoted father of immunology, Paul Ehrlich, coined the term “magic bullet,” medical science has marched towards more personalized drugs that target key molecules that cause diseases including cancer. 1 We find ourselves now, over a century later, in an exciting era of discovery that has produced many antibody drug conjugates (ADC) designed to precisely target the diseased cells and not healthy cells. ADC uses this strategy to take advantage of the specificity of antibodies while delivering a covalently linked cytotoxic payload directly to diseased tissues to reduce the multitudes of side effects and toxicity. 2, 3 As basic research identifies more targets and antibody engineering procedures improve, the range of antitumor and anti-disease weapons may seem limitless.
I am immensely proud of being an alumnus of the University of California at Berkeley, where I was able to get a world class education and have opportunities to meet with and learn from superb professors, some of whom have since earned Nobel Prizes. Those were some of the most fun years of my life and I also appreciated the beautiful, sprawling campus with lots of fantastic architecture and wide-open green spaces to lounge around on and play catch with my friend every now and then. It is mere coincidence that the day this article published is also Marian Koshland's birthday, and it got me thinking about Koshland Hall, one of the newer (now old, because so am I) buildings when I started college, and which Koshland it was actually named after.
Having worked in a proteomics lab for my PhD dissertation, I had some familiarity with the tools and strategies used to study biology on a systems level. One of the concepts I was always interested in was the ability to just follow a protein's journey throughout the cell, from the time it is translated by the ribosome to its final destination either within an organelle or when it is secreted into the extracellular space. At the time I was finishing up, I wasn't sure that the technology was yet advanced enough to make that a reality, particularly if done within a single cell. But within the past few years, a new era of spatial proteomics has emerged to allow us to observe cell biology in a whole new light.
Just a few short weeks after the highly irreverent yet still important Ig Nobel Ceremony, the science community recognized the cream of its crop with the 2023 Nobel Prizes in the first full week of October. The dates for the official announcements are aligned with their usual order throughout the years, always announcing Physiology and Medicine first, then Physics, then Chemistry. The Nobel Committee will transition toward the Literature and Peace prizes to round out the week before Economics is announced on the following Monday. As usual, these prizes recognize a lifetime of work that has given the greatest benefit to humanity. Click the links to check out some of our picks for greatest Nobel science achievements as well a look at last year's Nobel winners, but here we go for this year's running tally of scientific legend.
I recall a time many moons ago when I first started my graduate journey at Duke. I was doing one of my final rotations before joining a thesis lab, and I was sitting in a lab meeting where the group was discussing a particular surface marker on immune cells. Apparently this marker (long since forgot which one) could be cleaved and the "shedding" effect led to normal immune function. So silly young me who didn't know asked, "So what happens if you can't cleave it?" At that point one of the research professors said, "Well that's a stupid question" but in a way that was more bemused than malicious, as it turns out that was the thesis project for the postdoc in the lab who was training me. Other than the part where I probably should have known that was her entire project for like six years, I had stumbled upon my first "stupid" question that actually led to tangible answers that contributed to our understanding of science. Not that I actually did the work here, mind you, but someone else also asked that question and decided to answer it for themselves. I've long since forgotten the mechanism or the phenotype of the mouse that couldn't shed that marker, but the core memory stuck with me and shaped the way I approached students and education, because while questions might seem dumb, they at least always make you think.
I always look forward to this time of year, even more so sometimes than the actual Nobel Prizes, because I want to see what new insights can be derived from the weird science that, as they say, first makes you laugh, then think. That's right, now we are at the 33rd First Annual Ig Nobel Prizes! Just like last year and the few years before, the Ig Nobel ceremony was conducted virtually while the pandemic is still not quelled to an extent that allowed the organizers (men and women of science, see?) to be comfortable enough to have hundreds of people packed into a raucous arena, so the paper airplane tosses and everything else was pre-taped and released online. This did not take away from the absurdity and the few laugh-out-loud moments that I (and probably hundreds of thousands of science enthusiasts tuning in from around the globe) had during the 90-minute event. I do wonder if some of these might supplant my personal top ten, but maybe not just yet. Now let's see what happened!
Having navigated the worst of a pandemic, it is amazing how just barely over a year since the first reports of SARS-CoV-2, a vaccine was developed. What might not be as well known is that the foundation for the COVID-19 vaccine (and a just-as-rapid quell of the spread, though obviously still be vigilant and take proper precautions) was laid out for decades with the study of mRNA-based vaccines. Although I still masked up regularly until early 2023, I was thrilled to have the opportunity to get the vaccine as soon as it was available in 2021, as my family and I sought to protect ourselves and others through a day or two of aches and nausea. The scientists and technicians at the various companies that conceived of and manufactured the vaccines that have been distributed across the planet are relatively anonymous but should be commended. August being National Immunization Awareness Month, let us learn a bit more about vaccination and the pioneers that offer us important protections that we should not take for granted.
By the time I returned to graduate school to complete my PhD, the entering class in my program was half women and a large portion of my instructors were also women. I was encouraged to have access to their perspective and philosophies on science, and thought it a far cry from a decade before when most of my undergraduate instructors were men. Unfortunately, regardless of who is doing the reporting, although women do make up a good proportion of health care workers and just under half of all life science positions in the workforce, women comprise less than 30% of all STEM workers in the world, and since they are nudged away from science throughout their lives and careers while earning fewer STEM degrees than men, particularly in non-physical science fields. This is demonstrably worse for non-Asian minorities as their representation in STEM fields is under 10% overall. While I am thrilled to have been influenced by so many talented women scientists and colleagues in my career, these numbers can and should be improved.
Left-handed folks aren't as rare as people with polydactyly or syndactyly (webbed hands or feet), but you may have noticed that there aren't as many left-handed people out there as right-handed people. Some of this is due to societal pressures, but I was intrigued to learn that much of it may have to do with genetics, and these genetic mechanisms may extend not just to which hand is dominant, but how the body plan is determined. Like many things in bioscience, it isn't always apparent that handedness and its genetic components are critical to other aspects of physiology, but as we will find out together, the genes associated with whether you use one hand more efficiently than the other can also influence developmental and neurological health!
I was recording a new BioChat (you should subscribe) recently with a professor at Harvard. We discussed gene therapy in passing for the disease he was studying, and one of the things that he brought up was the need to ensure that whatever therapy is designed has to be safe and effective. This of course is the promise and also the challenge of CRISPR-based research, in which the model is known but the targeting efficiency isn't where it needs to be in order to be of practical use in therapeutics. This is a big reason why the biomedical research world was abuzz with the recent announcement (and preprint) of a major discovery by scientists at the Broad Institute, where they characterized the mechanisms and the potential utility of a system similar to CRISPR in eukaryotes.
As science advances, one of the recent trends that continues to pay dividends is immunotherapy to fight cancers. In many cases, the strategy is to mobilize the immune system to attack tumor cells based on cancer-specific antigens expressed either by the tumor itself or within the tumor microenvironment, either by stimulating normal immune cell function to their new tumor target or by removing the suppression of the immune system that is characteristic of many cancers. The trick is to find a way to attack only the tumor and not normal tissues, and certainly not to somehow trigger autoimmunity.
Ever since the official start of spring, I have been sneezing repeatedly with watering eyes possibly due to the various types of pollen and random detritus in the air. No matter your location or the time of year, we are always swimming in potential allergens, and some of us react to these generally innocuous substances more intensely than others. As summer is often a time of heavy pollination, let's explore allergies and what can be done about them.
Ever since my wife started listening to some true crime and fantasy podcasts a few years ago, I ventured on a different path with my podcast journey as I steered toward celebrity interviews, comedy, and the occasional science podcast. Of course you know we did start our own ABclonal podcast, BioChat, and previously we also highlighted a few fun and interesting science-themed podcasts that won't take too much out of your day as you commute or go about your work. In addition to entertaining and educating you, podcasts also have some mental and personal benefits. There are only so many hours in your day with plenty of choices, so let's see what we can do to help with some constructive distraction!
As a lifelong Star Trek fan, it has been exciting to see a lot of the science fiction gradually become science fact, even from the classic episodes with Captain Kirk and Mister Spock. From automatic doors to cellular phones, and even the computing innovations we take for granted such as the Google search engine, touch screen iPads, and the Alexa voice-activated assistant, science fiction like Star Trek has fed our imaginations to turn concepts into reality, such as this happy goofball (albeit a very resourceful goofball) making Doctor Octopus tentacles. I will continue sprinkling in Star Trek references because many of the neuroscience-based innovations in this post seem inspired by mere words on a script page that turned into an "aha" moment on screen, but as paraphrased from Arthur C. Clarke's laws, nothing is truly impossible with the right kind of science.
I recall having to make hundreds of custom peptides, and even though we used an automated peptide synthesizer rather than doing it by hand like another lab in the building, making a dozen peptides at a time still took hours on the machine and then another couple days for the purification and lyophilization processes. My mentor and I briefly joked about using bacteria and a polycistronic construct to just have the little guys use their ribosomes to do it for us, but the purification process probably would have been impractical. This does make me appreciate the wonders of natural biosynthetic machinery, and I promise there is a point here because we still use animal hosts to initially produce antibodies. Today, let's explore the process by which most commercial and custom antibodies are still produced, including here at ABclonal!
We live now in a polarizing environment where many people can't agree on issues that may seem obvious for the greater good, and part of that is likely due to a mistrust of scientists depending on one's education level and political leanings (probably the most diplomatic way I can phrase this). Science has brought us many wonders, from faster transportation, to lifesaving medicines, to the devices you are using to read this right now. Science also works to continue building our knowledge base, and perhaps one of the greatest examples of this is the banning of leaded gasoline, highlighted in an amazing episode of Cosmos hosted by Neil deGrasse Tyson. Because proper regulation is needed to keep both society and science in check, I thought today we would explore how science works with the law to ensure a brighter future for humanity.
May 22 is the annual International Day for Biological Diversity, which seeks to maintain and improve the diversity of plant, animal, and microorganism species across our planet through the protection and maintenance of their native ecosystems. Given the human dependence on biodiversity to sustain us, particularly in our food supply, this is an important issue to raise awareness for and to enact policies and mechanisms to provide food security and prevent famine and disease. With plants being the backbone of nearly every ecosystem and food chain on Earth, many scientists are considering plant biology to address these major global issues, especially as biodiversity continues to be threatened by global climate change and other human interventions.
Computers are ubiquitous in our lives now, particularly the majority of us who have a miniature supercomputer in the palm of our hands. With more efficient and powerful computer technology coupled with the understanding that biology is much more complex than dissecting out the role of a single protein in a signaling pathway, the relatively new field of computational biology and bioinformatics has exploded over the past few decades as researchers needed new tools and strategies to understand biology on a systems level. This has allowed non-traditional professionals to enter bioscience research, from primarily computer scientists to bench scientists who have taught themselves coding and statistics. As the computational component has permeated through nearly all of modern biology, we realize that there is a beneficial coexistence between the experimentalists and the keyboard warriors who make sense of growing datasets.
I'm a big proponent of telling fun stories, particularly about science (and sometimes baseball), and in just plain having fun, which I think is a good way to go through a life that is all too short on the geological scale. I also like staying in touch with my friends and making new friends, because a life well lived includes all the fine folks we've met along the way. As you may have seen in our other communications, we are getting set to launch a new podcast series we have named "BioChat," because we are going to be talking with a lot of folks who are associated with bioscience research and work in service of promoting better health, stronger research and clinical outcomes, and ensuring improved quality of life.
There was a fun seminar with a guest speaker once upon a time when I was in graduate school where he put the physiological effects of stress into perspective. I can't remember his name now, but the gist was that if you imagine yourself as an animal fighting for survival, you are very likely to want to keep the body ramped up to run or fight rather than to fight infection, digest, or reproduce. In a human context, when the person is under pressure due to a huge work project or some other external force derived from a crushing mountain of responsibility, they are thrust into a situation where the body and mind think that they are in a "fight or flight" mode and that can lead to many physical and psychological detriments. Although what we do in science is important and requires lots of work, it is important to realize that good science is harder to do if we are overcome with stress, so let's find some ways to manage stress so we can be more productive and healthy.
Apr 12, 2023 12:00:00 PM by Kin Leung
ABclonal in Action: Diabetes, Insulin, and Resisting Graft Rejection
As a scientist (or at least, a guy who likes science and works for a bioscience-oriented company), I am invested in the power of scientific research in improving the quality of human life across all arenas. It is particularly gratifying when our customers, who are all primarily research scientists, derive direct benefit from ABclonal's products in their published research. In this case, our business development director had a customer recently defend her thesis based on a publication that used several ABclonal catalog antibodies. I enjoyed reading her group's article that may lead to more effective treatment strategies for diabetes patients going forward, so let's get to it.
At the turn of the millenium, actor Michael J. Fox, whom you might remember from the Back to the Future films and Family Ties if you're a certain age, had to depart the cast of the popular sitcom Spin City because he was diagnosed with Parkinson's Disease. I was a frequent viewer of the show (and of course I watched Back to the Future, and so should you) so it was a shock to the system to see a man who was still so vibrant and young have to take a step back from his profession because of that diagnosis. April is Parkinson's Awareness Month, which you can read more about from Fox's foundation and others, and in this post I thought we could learn more about this disorder together as humanity works towards an eventual cure.
It has gotten harder as I've gotten older to keep the weight off, and I miss the days when I could eat all I could eat at buffets and not gain a single pound. I'm still not that big, but definitely have a bit of the #DadBod and my HbA1C count suggests that I'm bordering on prediabetic even as my cholesterol levels stay within established norms. It's a sobering reminder that as we age, the body simply can't be as efficient as it used to be, and even for those folks still far from "middle age," it makes sense to set up good healthy habits that will persist throughout however much time we have left on this blue marble flying through space. Being National Nutrition Month at the time of this writing, now is as good a time as any to reset reasonable dietary goals without having to go full cold turkey.
Everywhere I've been in school or at work, there has been at least one session about ethics, whether it was a semester long course, a small retreat, or just a statement in passing for the company to cover their legal obligations. I'd like to think of myself as someone who wants the best for everyone he encounters, and try to live my life based on acceptance and collaboration so we can achieve common goals for the greater good. Because people come from diverse backgrounds with different upbringings, it is hard to distill ethics and values into a simple set of parameters, but I also think that in general, most people know what is right and what is wrong.
Remember once upon a time when I said my first actual laboratory research project involved myelin basic protein? Other than knowing that the mother of one of my high school friends had been diagnosed with it, this was the first real exposure I had with multiple sclerosis. I eventually learned more about the immune system and autoimmunity, and the thought of your own body attacking your literal nerve cells was scary and made me feel for the people who have to live with and manage this disease every day. March happens to be Multiple Sclerosis Awareness Month, which as the name suggests works to make sure the public knows about multiple sclerosis, develops empathy and understanding for afflicted individuals, and encourages participation in events and activities to spread awareness. In this blog, let's explore the disease, current treatment strategies and ongoing research, and ways that you can help both in and out of the lab.
Previously, I listed some of my favorite educational YouTube channels, which includes the Kurzgesagt channel with its fun animations and soothing narration of scientific concepts, including many topics in biology and biological functions. As part of my algorithm, this one popped up in my feed recently:
As we head out of February and into March, we celebrated the resilience of our colleagues from diverse backgrounds during Black History Month and continue to promote the accessibility to science and STEM careers. There were a couple really important dates to recognize girls and women in February as well, with the National Girls & Women in Sports Day on the first day of the month, followed by the International Day of Women and Girls in Science on February 11. And now in March, we have the entire month to celebrate the contributions of women throughout our history. To kick off Women's History Month, I wanted to take a look at some of the most accomplished women scientists that advanced our knowledge to new heights while overcoming societal pressures and prejudice.
The last day of February marks a hopeful end to winter and a transition in the sporting world as well, but is also important in society as a day to recognize and raise awareness for rare diseases. Known as Rare Disease Day, the goal is this observance is to remind humanity that just because a disease is not prevalent or has as much research dedicated to it does not make it any less important, as certain individuals obviously suffer from these rare diseases and deserve accessibility to treatment and hopefully a cure. While it makes sense that more funding is funneled to cancer and neurology research since it affects so many more people, performing research in these comparatively uncommon maladies could offer insight into their diseases that get most of the research dollars.
When living in Chicago, we would often get off on the 31st Street exit on Lakeshore Drive, and at the time there was a sign pointing out some parking areas for McCormick Place. I recall specifically that two of those were labeled "E2" and "E3" and being the gigantic nerd that I am, thought immediately of ubiquitin. Since it is found in every type of cell in all eukaryotic cells, the protein is appropriately named. Check out the cool ribbon model I made of ubiquitin for my students once upon a time, and then let's take a look at ubiquitin's functions in living organisms and how it contributes to health and disease.
As I march slowly toward the twilight of my life, ever more I wake up with aches and pains and can hear the sounds of popping bubble wrap or Rice Krispies drowning in milk every time I make any major movements. Everyone deals with the realization of their own mortality in different ways. Some decide to finally climb Mount Everest or go skydiving. For me, I decided to look into the research behind cellular aging, and how we can make the most of our later years with the power of knowledge and biomedical science. This does remind me of that one episode of Star Trek where Jake and Nog have to get stuff for a mad scientist's cellular regeneration and entertainment chamber, which is supposed to restore the cells to a younger state and keep them from being literally bored to death. If you consider some of the treatments and technology being implemented or proposed these days, it almost seems like Star Trek has inspired yet another advancement beyond just cell phones and Alexa.
We’ve all been there…the experiment didn’t work for the 87th time, and the feeling of dread and impostor syndrome surrounds the mind as the seminar or thesis committee meeting looms. It’s easy to dwell on everything that isn’t working right at that moment, but there are scientific discoveries to be made, and someone has to do it, and it might as well be you! That’s easier said than done, but with the proper support mechanisms and some confidence boosts, your goals will be achievable.
Jan 25, 2023 12:00:00 PM by Kin Leung
8 Educational and Entertaining YouTube Channels for Your Down Time!
Like many of you out there, I used to pass the incubation periods for my experiments by scouring YouTube for videos, both of the educational and entertaining variety. While the bulk of this was admittedly cat and animal videos and human misfortunes (that normally did not result in debilitating injuries or death, mind you), I did prefer the channels that were a combination of educational and entertaining, which is a philosophy I adopted as a teacher and mentor. So today, I'd like to share some of my favorite channels that might help you de-stress from a hard day at work, and also probably teach you something! You can click the headers to go to their main channel as well as check out the example videos.
In the holiday rush, there were some fun science stories I was unable to get to other than the 2022 breakthrough of the year celebrating the ongoing JWST expedition. Now that we're back from celebrating with friends and family, let's check out some of what we missed!
One of the first things my K-12 science teachers taught me was to write a formal laboratory report. While comparatively crude, the basic structure of the middle/high school lab report is similar to what you might see in an article you read in Cell or Science. I've helped write numerous grant applications and journal articles as well as my own dissertation, and I even helped teach a course at the University of Chicago that emphasized grant writing, so I've been intimately familiar with how the process goes. Additionally, as various lecturers have told me, a good scientist is also a good storyteller, so it made sense that their message stayed with me all these years because they told very good stories about their work that were embedded into my memories (and my memory is honestly not that good). I thought I'd take this time to help you all develop your writing structure and consider how best to deliver your message, whether it is in your own grant application, a new journal article submission to present your findings to the world, or a speaking engagement that, well, keeps the audience of your very well-informed peers engaged!
Most of the United States is feeling the cold of winter at the moment, but it's never too early to start planning your compost setup. As you all may guess, I'm a big proponent of sustainable living, even in the laboratory, and the habits we develop could also lead to a healthier yard and surrounding natural landscape. As a teacher in Chicago, I developed an engineering project with my students to turn a part of our school courtyard into a vegetable garden, and we incorporated composting into it. It was a very cost-effective project as the students would bring in their unused vegetable scraps, recycled papers, and egg shells to school and develop their own compost mixes from which we would derive fertilizing materials for corn and squash. We ended up growing quite a bit of corn and not much else, but I anticipate that was because of the suboptimal lighting due to our building blocking the bulk of the sun's rays most of the day. But imagine what you could do with a little planning and a bit more budget than a public school (but that's a story for another day)!
You know how it is this time of year, when you have to make some New Year’s resolutions to improve from this past year. The good news is that you can put off the actual execution of said resolutions until next year (since it’s only a few days away), and it is probably true that you won’t be able to keep some of them, but at least you can say you tried! Writing these down is a good way to keep a record and try to keep yourself accountable. Besides the normal ones about more exercise and losing weight, here are some science-associated suggestions for good resolutions that aren’t that hard to keep!
With the holiday season upon us, it is a time to relax and be among loved ones again as we recharge for the push into the new year. Along the way, we have learned quite a bit about medical advances and new discoveries into life processes, information that will be used to drive the next stages of innovation. One of the great perks of having been in an academic setting was the constant immersion in ideas and collaboration, but even though some of us are no longer connected directly to academia, the vast interconnectivity provided by the internet means that we are never too far away from good information that could teach us something new and exciting.
Every December, Science Magazine awards a scientific breakthrough of the year. When you take a look at the previous winning breakthroughs, they come from all different fields and many have been eventually awarded with Nobel Prizes. The breakthrough from last year, for example, is particularly important for structural biologists who hope to translate their findings into practical applications for other biologists and drug researchers. Since a breakthrough suggests a more recent discovery, it is no surprise that the Science Breakthrough of 2022 is the NASA JWST that has brought us myriad breath-taking images over the past year since its launch.
Source: NASA
What is STAT5B?
STAT5B belongs to the STAT (signal transducer and activator of transcription) protein family, a group of latent cytosolic transcription factors activated by Janus kinase (JAK) tyrosine kinases. The JAK-STAT signaling pathway is responsible for many important biological processes including cell proliferation, differentiation, apoptosis, and is also involved in the modulation of a variety of cytokines to control the immune response.
December is a month of holidays and celebration, but it is also a time to raise awareness for a global epidemic that has lasted over four decades. During World HIV/AIDS Awareness Month, health organizations, including the United States Department of Veteran Affairs, serve to remind everyone about the importance of getting tested, to remember those who succumbed to the disease, and to improve access to advanced therapies.
Since its first identification and description in 1981, medical advances have offered effective therapies to keep the virus at bay, and in some cases even completely cure a patient of the human immunodeficiency virus, or HIV, and to prevent it from becoming the acquired immunodeficiency syndrome, or AIDS, which is often catastrophic to the patient. Unfortunately, as of 2021 per the World Health Organization (WHO), there are still over 38 million people living with HIV, with approximately 1.5 million new infections and 650,000 HIV-related deaths. Much of this has to do with lack of education or proper infrastructure and often obstacles to accessibility for treatment and prevention. I hope to explore HIV with you during this month of awareness so we can do our part to mitigate this persistent epidemic.
It has been a very trying time for all of us across the planet, with COVID-19 still lurking around as new variants pop up, while having to also deal with the growing spread of monkeypox. As for me, I’ve had a couple of bad colds since the start of the pandemic, but one was before commercial testing was available (maybe COVID? But probably not!), and the other was more recent and was definitely not COVID (lucky me!). In fact, I have to say that because of certain choices I have made to avoid the big bad disease as well as other preventable diseases, this has been the least I’ve experienced illness in quite some time.
It’s that time of the year again where we’re supposed to gather with family and close friends, talk about anything but politics, and eat monstrous amounts of good food throughout the next few days as we celebrate Thanksgiving. I used to joke with my friends that we would all be subjected to tryptophan poisoning, but just as the story about Ben Franklin wanting the national bird to be a turkey is just a myth, we can’t blame our post-feast stupor on just tryptophan either.
Nov 16, 2022 12:00:00 PM by Kin Leung
Trim and Proper: A Nifty Method for Targeted Protein Degradation
A common experimental strategy in studying the effects of a specific protein in cells or organisms is to remove it. One can determine the physiological outcomes in the absence of that protein to ascertain its relative importance in maintaining normal functions, or in some cases, to note that it is dispensable or redundant and might have a backup within the cell to take up the slack. Some targeted techniques include RNA interference (RNAi) and CRISPR-based gene editing, and in many cases, it is possible to generate knockout cell lines or even organisms, like mice, that cannot express a specific protein. But when those strategies are not feasible for the experiment at hand, what is one to do?
My wife’s family has a rich and decorated military history, with her grandfathers, in particular, having served honorably in major combat. When I was in graduate school, I had a few acquaintances who had served in the Navy or Marine Corps prior to using their military college benefit to go into the sciences and eventually earn their doctorates. The military and those who serve are a key cog in our society to help protect the many freedoms we enjoy, and while they’re not as well-known as, for example, athletes like Ted Williams who served in combat, scientists have contributed greatly to efforts in war and in peace.
Oxidative stress is an imbalance between the production of reactive oxygen species (free radicals) and antioxidant defenses. [1] The body’s cells produce free radicals, which are nitrogen- or oxygen-containing molecules with an uneven number of electrons [2], during normal metabolic processes. [3] Meanwhile, cells also produce antioxidants that neutralize these free radicals to prevent excessive cell and tissue damage. In general, the body is able to maintain a balance between antioxidants and free radicals. [3] However, this balance could be disrupted under certain conditions or environmental stress or infection, and uncontrolled oxidative stress can accelerate the aging process.[3]
Slogging through another election cycle, you are probably deluged with dozens of political ads every day with candidates touting what they will vaguely do for your vote, and often, talking about how much their opponents suck. Don’t you wish they would have more concrete plans, or at least promise to delegate to true experts who want a better quality of life for the citizenry? As you determine which option to go with at the ballot box, perhaps it is time to consider what their position is on important science policy.
Oct 26, 2022 12:00:00 PM by Kin Leung
Vetting Your Sources: Confirming the Veracity of Reports and Data
Throughout graduate school and even now, I’ve relied on Wikipedia as a valuable resource for quick information. My mentors and teachers have cautioned me against actually citing Wikipedia articles, but often these articles will show up as top searches on Google, and their listed references lead to published scientific articles so I could always go back to the original source and see the data and conclusions for myself. The fact that Wikipedia is free and freely edited makes it prone to fictionalization, which reminds us of the importance of corroborating whatever we read with third-party sources and our own experiences.
The term apoptosis was first used in 1972 to describe a morphologically distinct form of cell death. Since those early experiments and observations, apoptosis has become one of the focal points for biological research, with myriad laboratories and research groups continuing to work to further elucidate the components and pathways that drive this programmed cell death. As a fundamental biological pathway, apoptosis has benefits and adverse effects for the host organism. For example, many therapeutic strategies involve the activation of apoptosis to kill cancer cells, while other treatments seek to prevent apoptosis to preserve precious cells in key tissues.
As human beings with trillions of cells, each of which has their associated millions of copies of myriad proteins and other biological molecules, it’s something of a miracle that enough of the molecules bump together at the right times to keep us alive and functional. In addition to our own cells, we also coexist with microscopic neighbors, including various beneficial bacteria, while fending off pathogens like disease-causing bacteria, viruses, protozoans, and fungi. We often consider the bacteria and viruses in most human diseases, which invoke our immune systems to fight them to keep us healthy, but it also makes sense that the fungi can affect us as well, a topic in cancer research that is gaining attention.
Many of you are well on your way through graduate school, itching to earn that precious PhD, while some are just starting out, getting ready to take your first midterms while preparing to choose your first research rotations. Regardless of where you are in your career, or even if you've already earned that doctorate and are on your way to a postdoc and beyond, ABclonal's blog series has put together some articles that can help you get through the day. Whether it is experimental troubleshooting or just trying to get along with your lab mates and PhD supervisor, here is a collection of previous blogs that should be of use to you.
Reaching the golden years doesn’t always feel so golden. As we age, disease, injury, and other stress factors from the environment will damage our bodies' cells. Most cells may be able to repair that damage, while our immune system usually clears those damaged cells through a process called apoptosis.[1] However, if cellular repair and clearance is not effective, the residual damaged cells will further weaken the immune system and deteriorate other biological processes. Is there a possibility that we can avoid this cellular damage and improve the health of older people? A cellular state known as senescence might hold the key to this question.[1, 2] During senescence, the damaged cells irreversibly stop dividing and resist being removed. [3] Researchers have shown that determining senescence biomarkers could lead to new therapies for the inflammatory disease caused by senescence in older people.[4]
As we try to come back to some level of normalcy after a couple of long, stressful years of pandemic, science has been continuing to chug along to improve the human condition. In celebration of this, we had silly achievements that made us laugh, then think, in the form of the Ig Nobel prizes, and this week, the cream of the crop was recognized with the three science Nobel Prizes. We wanted to highlight the Physiology and Medicine prize separately since ABclonal is a bioscience reagents company, but as we said before, every field of science is important to the pursuit of not just biological advancement, but the betterment of all humanity. So while you can also read about the achievements of the Medicine winner, Dr. Svante Pääbo, in the previous entry, here are the science prize winners in all the glory we can give them in this blog space!
Oct 3, 2022 6:59:46 AM by Kin Leung
2022 Nobel Prize in Physiology/Medicine Celebrates Human Evolution!
I had anticipated that 2022's Nobel Prize might go to something more contemporary, like the RNA-based vaccine technology or even the malaria vaccine, but as is sometimes the case with the Nobel committee, this year they threw us a pleasantly surprising curveball with the prize in Physiology or Medicine. With this award, the Nobel Prize definitely awards someone who gave the greatest benefit to humankind indeed as the recognition was for discoveries that look into the very origins of humanity!
Every fall, the world comes to attention for the unveiling of the Nobel Prizes, considered the most prestigious awards for achievements in science and the humanities. Per Alfred Nobel’s will, the original five prizes were to be awarded “for the greatest benefit to humankind,” and in 1969, the Economics prize was added to the mix.
That time of year that we've all been anticipating is here! That's right, the 32nd First Annual Ig Nobel Prize ceremony took place on the evening of Thursday, September 15, and the 2022 winners took their bows and hammed it up in one of the most favorite of scientific gatherings. Alas, the ceremony was online-only due to the COVID-19 pandemic, but that did not take away from the fun and love of science that is expected from this festival of glee that features actual Nobel Prize winners! Without further ado, please read on for the many great, uh, achievements by this year's newest additions to the Ig Nobel ranks.
Science is the process that allows humans to identify a problem and devise experiments to examine that problem and provide solutions. Not all scientific problems are created equal, but we can agree that all science, even those super mundane details that barely anyone ever thinks of, can be important to the whole of human knowledge. One of the great draws of science is that it is both rewarding and fun, and if ever there was a repository of high-brow humor, science is it. While we celebrate great discoveries every year with the Nobel Prizes, we also have an uproariously entertaining time with the annual Ig Nobel ceremony, a somewhat obscure but highly-anticipated event for the entire scientific community that shows the fun and human side of the "lauded" researchers. Let’s look at the Ig Nobel Prize, and some of the “greatest” Ig Nobel recognitions since the award’s inception in 1991.
Sep 7, 2022 12:00:00 PM by Kin Leung
Why Accessibility and Collaboration are Critical to Good Science
Like most PhD students, I had to generate publications in order to earn my doctorate, and my mentor had to use those publications to support his grant applications. The ability to churn out publications in volume is critical to sustaining academic research, and the labs with the best reputations are the ones who regularly crank out quality articles that land in top-tier journals. However, does science have to be driven this way? Are we overlooking some great scientists or discoveries because they were unable to find the opportunities for funding and exposure, and thus dropped academia in search of another career? We should explore improving accessibility to funding and publication in research so that everyone has the chance to share their ideas.
You may have stumbled upon many articles about the poor whales swallowing tons of plastic waste, flooding that is affecting communities and even national parks, or chemicals that are constantly threatening marine life. As the global temperature continues to increase, the ocean levels gradually rise, and life as we know it is threatened, it is almost like we are on our way to the apocalypse. However, humans do not have to accept this doomsday scenario! There are many things beyond our control that we will have to persuade our elected leaders to drastically change policies to conserve our natural resources, reduce pollution, and preserve biodiversity. But there are also many other things well within our control that we can do in the lab and at home to make a difference, since small actions will add up to significant positive change.
If you are reading this, either having earned your first faculty position or about to embark on leading a huge project, congratulations! You have obviously demonstrated the creative problem solving and other skills needed to successfully carry out and complete a scientific study…but maybe you’re not confident in your ability to lead or mentor? I would argue that many experiences you have accumulated up to this point will help you become the best mentor you can be, so let’s get to it as you cultivate the next generation of great researchers!
The arguably most fun thing about science is when your supervisor tells you to just do Experiment X to test hypothesis, but then they kind of forget to tell you how complicated the techniques are to perform that experiment, not to mention all the optimization you would need to do. I personally have never done a chromatin immunoprecipitation (ChIP), and since I wasn’t in genomics, the most sequencing I ever did was setting up quick reactions for the core facility to tell me that my gene constructs were correctly built. ChIP does sound rather simple when explained in class, but when you read up on the protocols,1 there are some limitations to what ChIP can do, especially given the large amount of starting material you need for the typical experiment. Luckily, in recent years, scientists have started to use an alternative technique called Cleavage Under Targets and Tagmentation, or CUT&Tag, which ABclonal is pleased to support through our antibody reagents.
When I was in college, I enjoyed reading about Chindogu, which literally means “weird tool” in Japanese. The whole point of Chindogu was to make hilariously “unuseless” objects, somewhat like a tool that you might use, but wouldn’t actually buy because it was so absurd. An example of such absurdity is this Hay Fever Hat, and there are countless others that I would recommend you read and laugh about. Although Chindogu are essentially impractical devices meant for laughs, I got to thinking about how I MacGyver’d through graduate school in repurposing equipment and designing new ways to make my lab life easier even as our funding dwindled. Known affectionately as lab hacks, I’m sure you can find some of these on the internet, but I’ll share some of my favorites here.
Since I’ve been living with it for as long as I can recall, I don’t consider my visual impairment a disability. Unlike the millions of people who require corrective lenses, though, my impairment is much more permanent and far less manageable, but it hasn’t prevented me from enjoying life and participating in physical activities. I thought I’d take this time to talk a bit more about most genetic disorders that affect vision, and what is being done to achieve a better understanding to try to reverse the vision loss.
Jul 29, 2022 12:00:00 PM by Kin Leung
5 Steps to a Better PCR: A Troubleshooting and Optimization Guide
Ever since Kary Mullis (that crazy guy, may he rest in peace) officially invented the polymerase chain reaction (PCR), an entire generation of molecular biology has exploded across the globe as scientists use PCR for a number of applications, from measuring gene expression to forensics. While the textbook technique is relatively simple, as I (and many other fellow researchers) can attest to from experience, producing an ideal PCR is far more challenging due to multiple factors.
Jul 27, 2022 11:29:58 AM by Gavin Zhang
The ABclonal Advantage: Working With an Original Manufacturer
When you consider which of the dozens of biological reagents companies to work with, how can you determine which one is the right fit? There is, of course, a business aspect to making and distributing quality antibody reagents. The source of the antibodies that you rely on for your research will matter in terms of supply consistency, lead time, cost, and the associated services to support your product.
Jul 26, 2022 10:59:14 AM by Kin Leung
Potential Fraud and the Need For Vigilance in Scientific Review
I will admit that I am not a neuroscientist, having focused my research on immunology and cancer cell biology, but I’ve always been aware of Alzheimer’s Disease and the quest for better treatments and an eventual cure. It is because I am not a neuroscientist that I rely on the word of purported experts in the field who have dedicated their careers to finding these answers. There are various caveats like the level of journal the research is published in, the quality of the images (at least to the naked eye), the number of times the research is cited, and the known reputation of the authors, that help to determine the level of trust one can put into the finding. Yet, we find that some things still might slip through the cracks, and this reminds us that we need to scrutinize data more thoroughly to hold each other accountable and maintain trust in science.
In another life, I taught high school biology and had a lot of fun doing it. I had my students do the Cell City when we worked with organelles in the cell, and once we got to the genetics unit, we did something fun called Dragon Genetics. In this activity, students would pair up (one was the mommy dragon, the other the daddy dragon) and throw “chromosome” sticks to see what traits they would “pass on” to their theoretical dragon baby. The activity is quite simple once students understood basic Mendelian genetics (and some of the non-Mendelian patterns as well), and even my son was able to draw his own dragon baby when I had him be my guinea pig while he was still in elementary school. (Figure 1) There were some amazingly creative dragons adorning my classroom, and I hope you can share the Dragon Genetics activity with any teacher friends as we discuss non-Mendelian traits and disease here. As we celebrate the beautifully-designed experiments by Gregor Mendel that led to the modern study of genetics and genomics, we might also be reminded that patterns of inheritance, like many things in life, are far from binary.
Jul 18, 2022 11:59:58 AM by Kin Leung
What to Think About Zinc: An Essential Element for Healthy Living
Perhaps we only think of zinc as the extra element in our coins to keep manufacturing costs down, or as that random clip from the Simpsons about a world without zinc. Aside from thinking it is a wacky sounding word (I did look up the etymology and it is rather appropriate!), we just don’t consider zinc as being all that important. Once the pandemic hit, though, I noted that Costco was marketing their zinc supplements a lot more, and after doing some extra research, I bought some to add to my diet.
Jul 13, 2022 12:00:00 PM by Kin Leung
ABclonal in Action: 10 Scientific Studies Using ABclonal Antibodies
Open collaboration is important for sustainable science, and every new study or publication, no matter the journal or institution, contributes to a greater understanding of biology, for better or for worse. Dozens of prior discoveries funnel into every new breakthrough, so we need to appreciate the years of painstaking labor and thought that go into every new morsel of knowledge. It is very fulfilling when ABclonal products are part of the fuel that drives these studies in diverse fields of biology. With our ABclonal in Action series, we hope to highlight our products as well as the new insights from our customers all over the globe that will become stepping stones for the next generation of cutting-edge bioscience.
You go through everyday life thanks to the intricate communication and interaction of tissue and organ functions between the trillions of cells in your body. Within those tissues, a non-cellular component exists called the extracellular matrix (ECM). Imagine a structure made of water, proteins, and polysaccharides that helps to give structural support to surrounding cells as a connective tissue. Within the ECM lies a group of enzymes named matrix metalloproteinases (MMPs). As endopeptidases, which are enzymes that break peptide bonds, the main role of MMPs is to break down collagen and other proteins in the ECM, whether in normal tissues or in promoting cancer metastasis. MMPs are divided into collagenases, gelatinases, stromelysins, matrilysins, and membrane-type (MT) MMPs, as well as some other non-classified MMPs.[1]
My wife and I used to watch House, M.D. starring Hugh Laurie, in which he was a cranky doctor who happened to be a Holmesian genius in diagnosing rare or mysterious diseases. We are fortunate to have doctors who have much better bedside manner, but as an entertainment option, House was a lot of fun. One of the running gags for fans of the show is that the mystery disease of the week is never lupus, except for the one and only time that it was. My fond memories of this show got me to thinking about how difficult it is to diagnose lupus, and about other autoimmune diseases that still remain mysterious and challenging to treat. I decided to find out how modern medicine is approaching this continuing health issue.
Jul 1, 2022 10:50:18 AM by Kin Leung
Controlling Monkeypox Spread: Protecting Against the Next Pandemic
It seems like decades since the SARS-CoV-2 pandemic shut down the world economically and socially, and even now we are not fully out of the woods. The COVID-19 coronavirus continues to persist, hovering on the cusp of becoming an endemic disease after having caused over one million deaths in the United States alone out of over six million deaths worldwide since the first reported cases in 2019. Although the various coronavirus vaccines have conferred some level of herd immunity across the globe, the danger of mutations causing variants that might escape vaccine protection is real, so continued vigilance and best practices are key to returning to normalcy. Perhaps our resolve as a global community and as a species will be tested in short order as the monkeypox outbreaks surge.
Once upon a time when I was a fledgling science nerd in high school, I started learning about the process of apoptosis, which remains to this day the most studied form of cell death in various functions including organismal development and defense against cancer. As an immunologist-in-training, I also learned about the classical complement pathway that the immune system uses to destroy infected cells, and also necrotic cell death or necroptosis (which is full of really gross pictures if you dare to Google it). Of course, I learned about autophagy in graduate school and really appreciate its utility in normal physiology and disease, while very recently I read about ferroptosis as yet another programmed cell death (PCD) pathway. Right around when the Nobel Prize was awarded to recognize the elucidation of PCD, pyroptosis came about as a novel PCD pathway that is continuing to gain steam in its clinical relevance. It seems logical for cells and organisms to have redundant systems in place to clear away damaged and malignant cells before a health crisis can emerge if the cell evades the primary route of apoptosis.
My first experience in a basic research laboratory was a structural biology project, in which we were attempting to solve the structure of a nervous system protein known as myelin basic protein (MBP). As a rookie undergraduate scientist at the University of California, I had great mentors who taught me everything, from how to purify recombinant proteins from bacteria to doing library work to understand what had been done before so I could build upon it. I also learned how to use an electron microscope (EM) to gather structural data. MBP was an interesting challenge as it had multiple isoforms due to alternative splicing, and generally behaved like a random coil. 1 The major function of MBP is to take advantage of its highly positive charge to compact myelin in higher organisms, with research over the years suggesting it may have some capacity to form alpha helices, although atomic-resolution structures have not yet been reported. MBP has also been reported as a biomarker in autoimmune diseases such as multiple sclerosis. 1
Jun 17, 2022 12:00:00 PM by Kin Leung
A Path To Effective Precision Therapeutics For Alzheimer’s Disease
Before my grandmother passed, she had been battling severe dementia for a very long time, which made it difficult in many ways to have conversations with her. It would take several minutes for her to process who I was, and then it would seem like she would remember me and my family, but she would still have to ask for clarification several times even after we had answered her queries. I am grateful that she is in a better place now, but her challenges in the final years of her life deepened my empathy for people who suffer from dementia, and those who take care of them.
Every now and then when I get hungry, I joke that my stomach is about to digest itself. For the longest time, human science was unaware that our cells could literally eat itself (or more precisely, parts of itself) as well! First described in the 1960s by Christian de Duve (who won the Nobel Prize for discovering the lysosome), the term autophagy derives from Greek words combined to mean “self-eating” and describes a process by which the cell degrades large components and organelles in a distinct mechanism. 1-3 The phenomenon was not studied extensively until the 1990s, when Yoshinori Ohsumi performed a series of groundbreaking experiments to determine the underlying mechanisms of autophagy, an achievement for which he was awarded the 2016 Nobel Prize in Physiology or Medicine. Ohsumi’s work has led to an explosion of research that has precipitated a greater understanding of the role played by cellular digestion, degradation, and recycling pathways in human health and disease.
Jun 3, 2022 12:00:00 PM by Kin Leung
Traffic Management: The Indispensable Vesicular Transport System
When I taught high school biology, a favorite part of the curriculum was cellular structures and functions. I set up an activity suggested by other experienced biology teachers that was based on the “Cell City,” a learning analogy where students would create an artwork of a city with the mitochondrion as a power plant and a vacuole as a lake. (Figure 1) I wish I saved their very creative projects, but I distinctly remember one group used the Chicago Transit Authority’s elevated train system map to represent the endoplasmic reticulum (ER), a very clever use of the analogy and a nod to city pride. It was also the first time these students really thought about vesicular transport, although they didn't fully understand its importance.
When I was growing up in Hong Kong, and even after I came to the United States, my parents and grandparents would periodically give me ginseng beverages and soups, which was not always pleasant due to the bitter taste. As a result, I don’t think I really appreciated the benefits of ginseng, both scientifically confirmed and perceived. It is fun and informative to read about the myriad studies of natural plant extracts and how they can improve our well-being. Many folks like to drink herbal teas or use plant-derived supplements such as aloe vera lotions, so maybe this is good incentive to grow more of these beneficial plants such that they can provide health products as well as some clean oxygen for us to breathe!
With a background in both immunology and cancer biology, I’ve always had a fascination with the interplay between the body’s immune system and any tumors that might pop up. Originally, it made sense that the immune system would actively seek out and destroy cancerous cells, but the emerging consensus is that the interactions between cancers and host immunity is far more complex. In addition to growing new blood vessels and reprogramming metabolic processes, there appears to be some imbalance between avoiding immune cells while also promoting tumor-infiltrating inflammatory cells to promote its growth. 1 (Figure 1) Trying to dissect this apparent contradictory relationship between tumors and host immunity remains a hot topic.
May 20, 2022 12:00:00 PM by Kin Leung
No Rash Decisions: Novel Treatment For Genetic Skin Disorder (UPDATE)
Throughout the COVID-19 pandemic, I have been washing my hands with vigilance to prevent the spread of germs. As a result, the skin on my hands have become calloused on some parts and mostly dry, with cuts and slight bleeding on occasion. I thought this was inconvenient, but when I learned about children with a rare genetic skin disease, I stopped feeling sorry for myself and dug a bit deeper into their plight. After all, my skin issues are just due to excessive hand washing (which everyone should be doing anyway!); these poor kids have to live with this painful disease, known as dystrophic epidermolysis bullosa, for their entire lives.
Towards the end of my doctoral research, I first heard the rumblings of an acronym termed “CRISPR” that was starting to gather momentum. By the time I earned my doctorate, the applications that were discussed in both theory and in practice accelerated to the point that, while I didn’t fully understand the mechanism of the factors involved, I was certain that the discovery and re-engineering of this prokaryotic phenomenon would eventually be recognized with a Nobel Prize. Less than a decade after their first publications on the topic, 1, 2 Emmanuelle Charpentier and Jennifer Doudna were awarded the Nobel Prize in Chemistry “for the development of a method for genome editing,” which sounds a lot less important than it actually is!
Many of the most popular vacation destinations are in warm, sunny climates like Hawaii or Southern California, and there are larger human populations where people can actually go outside without having to put on a sweater. With the warm, comfortable weather comes exposure to the sun. Our sun, of course, is the center of the solar system, the constant supplier of natural energy on Earth, and at the same time, a dangerous source of ultraviolet (UV) radiation. While enjoying the warmth of the sun, we also need to protect ourselves from UV and the maladies it could cause.
In March 2022, the United States Senate approved the Sunshine Protection Act, which would make Daylight Savings Time (DST) permanent starting in November of 2023. There was still some healthy debate over whether Americans should accept Standard Time versus DST as their new permanent or keep the current system of “spring forward, fall back.” Regardless of whether we will have DST forever, there is broad consensus that the clock switch every March and November is disruptive to our sleep patterns and our circadian rhythms.
Whether to save energy, increase night-time Trick-or-Treat hours on Halloween, get those few extra minutes of sun to squeeze in the last innings of a Little League or high school baseball game, or just to normalize our sleep patterns, even a seemingly obscure issue like switching between standard time and DST is tied to our health and well-being in our society. And this is why we have to consider how sleep and the circadian rhythm can affect our physiology.
Apr 22, 2022 12:00:00 PM by Kin Leung
More Than a Feeling: The Science and Applications of Sensory Receptors
The 2021 Nobel Prize in Physiology of Medicine was awarded jointly to David Julius, of the University of California at San Francisco, and Ardem Patapoutian, a neuroscience researcher at the Scripps Research Institute in La Jolla, California. Working independently, Julius and Patapoutian discovered the key receptors (TRPV1, TRPM8, Piezo1, and Piezo2) in our bodies that sense heat, cold, and touch. Their work not only helps us to understand how we perceive and adapt to the surrounding world, but also paves the way for drug discoveries that target a wide range of diseases, including chronic pain, respiratory disease, and cancer.
Have you ever entered a lab where it looks like a disaster area? It may not have happened after an actual centrifuge accident or explosion, but you can tell that the lab needs a makeover in every sense of the word. Some cases are on the extreme end, such as this lab at Georgia Tech that was an unfortunate victim of negligence, but for the most part it may be just a messy neighbor who needs a gentle reminder to take a moment and clean up their bench for the greater good. (Figure 1)
Apr 8, 2022 12:00:00 PM by Allen Zheng
The Cytoskeleton: Its Functional Importance in Cancer Research
Cancer remains one of the most prevalent and deadly diseases affecting humanity. According to the Centers For Disease Control, cancer was the second leading cause of death in 2020 for Americans behind heart disease. The American Cancer Society projects at least 600,000 deaths due to cancer each year, despite the fact that mortality continues to decrease each year. The majority of these deaths are from advanced cancer, which are cancers that do not respond well to treatment and therefore cannot be cured. It is when the advanced cancer progresses to a point where it can escape the primary tumor site, a process known as metastasis, that the prognosis becomes grim.
When I was an aspiring (much younger) scientist, one of the challenges was finding quality antibodies to accommodate our research group’s high-throughput Western blotting platform 1 while studying signaling pathways in cancer cell lines. When I got into marketing, I learned about ABclonal’s high-quality, high-specificity, and high-affinity antibody products. I really wish that I had access to these products when I was doing my thesis research! With a team of passionate, capable scientists supporting these quality products, I was thrilled at the opportunity to be part of this company and to help spread ABclonal’s brand to the scientific community.
In November 2021 we hosted a Lab Member of the Year contest. This contest allowed for various members of the research community to be highlighted for their contributions to their respective labs. At the end of it all Dr. Francesc X. "Xavi" Ruiz Figueras, an assistant research professor at Rutgers' Center for Advanced Biotechnology and Medicine, took first place and became ABclonal's Lab Member of the Year.
Dr. Ruiz Figueras is a highly skilled scientist who has made significant scientific contributions to the fields of enzymology and structural biology studying the structure and function of human and viral proteins, especially HIV-1 reverse transcriptase and human aldo-keto reductases, with an emphasis on catalysis and inhibition for drug discovery1. In his spare time he enjoys playing basketball, reading, and spending time with his family. Currently, he is reading a book about philosophy, which he feels has applications to living during these pandemic times. He believes we should take whatever the life lessons we can take from this book in order to hopefully be more resilient.
We sat down with Dr. Ruiz to discuss his lab life at Rutgers, his current research, and what he is looking forward to researching in the future.
Jan 11, 2022 12:57:34 PM by Edward Li, Ph.D
The 3CLpro as a Potential Target for the Intervention of COVID-19
The coronavirus disease 2019 (COVID-19) pandemic caused by SARS-CoV-2 has raised global health concerns. As case numbers continue to climb, there is an urgent need for an active drugs against SARS-CoV-2. The development of new drugs is time-consuming and costly, and the safety of new drugs is paramount. Therefore, the strategy of drug repurposing represents one of the fastest approaches to have an active drug to fight SARS-CoV-2 during the COVID-19 pandemic.
As a matter of fact, in silico repurposing approaches have found increasing popularity during the COVID-19 epidemic [1], especially with the great breakthrough achieved using 3CLpro as a target to screen drugs. By the end of 2021, the FDA has authorized the first oral antiviral drug Paxlovid, produced by Pfizer, to treat COVID-19. Due to that much of the scientific and clinical work on drug repurposing or drug screening against SARS-CoV-2 or COVID-19 is still ongoing, in this blog we will review the latest progress on the potential targets, including 3CLpro, for the drug discovery or intervention of COVID-19.
The cell cycle is a series of phases that takes place in a cell as it grows and divides. The cell spends most of its time in interphase. During this interphase the cell grows, replicates its chromosomes, and prepares for cell division. Once the cell leaves interphase, it will undergo the process of mitoses and start divining in order to create daughter cells. These new daughter cells will then enter their own interphase and begin a new round of the cell cycle. The cell cycle and its cues are of the utmost importance, because without the cues the cells can either multiply continuously, forming masses, or will not multiply. These cues are cyclins which controls the cell cycle progression.
Dec 3, 2021 1:00:00 PM by Fanyun Fang
The Role of Tumor Microenvironments in Cancer Development & Treatment
The tumor is an abnormal tissue mass formed when cells divide and grow excessively within the body. Tumors can be benign (not cancerous) or malignant (cancerous). Benign tumors may become larger but do not spread to nearby tissue or other parts of the body. Malignant tumors, on the other hand, can spread nearby to tissue and can also be transmitted to other parts of the body through the blood and or lymphatic system.1 But we are no strangers to tumors and how the develop.
On the other hand, many of us aren’t as familiar with a tumor’s environment. Tumor progression is profoundly affected by the subtle interaction of tumor cells with immune and non-immune cells within their environment. In particular, the interactions with the immune cell component of a tumor are fundamental in determining whether primary tumors are eradicated, metastasized, or established by dormant micro metastases.3 The environment that a tumor grows in is also much more complex than one would think because of its highly variable cell composition, large number of proteins, and structures involved in tumor formation.
This being said, tumor microenvironment includes:
• Heterogeneous populations of cancer cells
• A variety of resident and osmotic host cells
• Secretion factors
• Extracellular matrix proteins
Interleukins are a group of small signaling molecules, and a type of cytokine. They play a vital role in the body’s immune response by activating and deactivating immune cells. Recently, interleukins have gained visibility as a target to help treat COVID-19, and the WHO has recommended giving IL-6 inhibitors to patients with severe cases. Additionally, because of its widespread impact on the body, the interleukin family has gained popularity as drug targets over the last few years.
CD antigens have played a significant role in both diagnosis and treatment for several diseases ranging from autoimmune diseases to cancer. CD antigens are often used as drug targets in drug discovery and as biomarkers in diagnosis because they are both highly specific and are located at the surface of the cells to target different to identify and investigate cell surface molecules.
Since DNA was first discovered by researchers, decades of work have been done to understand its importance as it is the code of life itself. While DNA is the cornerstone of life, it is not immune to damages, and as so it is vital for DNA to repair itself for normal cell function to be maintained. Though, DNA is not always able to repair itself and this leads to some diseases such as various cancers. Fortunately, DNA repair pathways are capable of being tools to provide therapies to combat these diseases.
Having been over a year since COVID-19 was officially declared a pandemic, we here at ABclonal have had the chance to compile and explore a wide range of information, resources, and breakthroughs related to the novel coronavirus through our blogs. Below, we've organized and selected five recommended pieces of reading related to COVID-19, covering topics including asymptomatic cases, the different detection methods (PCR vs. antibody testing), as well as a one-year-mark review of the impact of COVID-19.
Foreword
To start, put yourself in a hypothetical situation: you have an identical twin brother who was secretly transferred to another family when you were less than a year old. His new family was poor, and your family was rich and therefore the environment in which you grew up was much better than his. After 50 years, by chance, you both happen to meet and it turns out that you look quite different from one another and are in different states of health; he is short and is suffering from heart disease, while you are tall, healthy, and are training to run a marathon. So, what was it that made you two so different, in light of the fact that your genetic materials are completely identical?
As a cornerstone of the body’s immune response, antibodies can provide significant data to support scientists’ research. Antibodies are used in a multitude of applications in research, including but not limited to: western blot (WB), immunoprecipitation (IP), immunofluorescence (IF), immunohistochemistry (IHC), chromatin immunoprecipitation (ChIP), and flow cytometry (FC). If you're looking to learn more about the various types of antibodies, their differences (ie. rabbit vs. mouse), and their uses in research, we've got you covered below with a collection of curated blogs from our ABclonal Knowledge Base:
It has now been over a year since March 2020, when the World Health Organization determined that the growing spread of COVID-19 would be officially characterized as a global pandemic. The year that followed challenged humanity with a public health crisis on a scale that few could even imagine, with far-reaching social and economic impacts still being felt across the globe. And yet, this past year also brought out the best in many of us, with medical professionals diligently fighting the pandemic on the front lines and researchers collaborating to progress our global recovery. Owing to the relentless work of researchers making breakthroughs in vaccine research, it appears that the long road to recovery has finally begun.
As a dedicated supplier of research reagents, we're passionate about making sure that your experiments run as smoothly and efficiently as possible. From our ABclonal Knowledge Base, we've compiled a list of helpful articles that address common issues and difficulties that you may run into while conducting your lab experiments, including ELISAs, measuring cell proliferation, western blotting, and casting SDS-PAGE gels. If you're looking for advice, troubleshooting tips, or recommended procedures, we've got you covered:
Feb 1, 2021 3:53:38 PM by Dennis Miao
Transcriptional Regulation of Myogenic and Metal Homeostasis Genes
On January 12, 2021, we had the privilege of hosting Dr. Teresita Padilla-Benavides, an Assistant Professor of Molecular Biology and Biochemistry at Wesleyan University in Middletown, CT, to present our first webinar of the new year. Her webinar discussed her research on the differential mechanisms for transcriptional regulation of myogenic and metal homeostasis genes. If you missed the live session of the webinar, we’ve got you covered here with a link to a recording of the webinar, as well as a recap below:
For the second installment in our ABclonal Webinar Series, we had the privilege of inviting Dr. Clarke Gasper, our Business Development Scientist, to share his insights on the production and development of monoclonal antibodies for cancer therapy. If you were unable to attend the live session or would like to review some of Dr. Gasper’s key points, we’ve got you covered with a recap of his lecture below.
Nov 25, 2020 5:27:12 PM by Kashyap Gayathri
Necroptosis: The Inflammatory Counterpart of Good Ol’ Apoptosis
A Bird’s Eye View of Necroptosis
Necroptosis is a type of regulated necrotic death driven by defined molecular pathways. Regulated necrosis regulates programmed cell death. Necroptosis is at the center of the pathophysiology of several clinically-relevant disease states, including myocardial infarction and stroke, atherosclerosis, ischemia-reperfusion injury, pancreatitis, and inflammatory bowel disease. Necroptosis results in necrosis-like morphological changes, such as cell swelling, plasma membrane pore formation, and membrane rupture. It also requires co-activation of receptor-interacting protein (RIP) 1 and RIP3 kinases. Necrosome is a complex formed by RIP1, RIP3 and Fas-associated proteins with death domain (FADD). Several studies in the preclinical stage have demonstrated that targeting necrosome can have variable effects on progression of tumors, indicating that it is largely cell-type or context dependent.
Autophagy can be understood as ‘self-eating’. In simple terms, it is a vitally important cleansing mechanism carried out by the cells in our body. It brings about the degradation of the cytoplasmic contents within membrane bound vesicles called lysosomes.
Nov 12, 2020 1:00:00 PM by Bryent Lee
Ferroptosis as a New Type of Inflammatory Programmed Cell Death
When it comes to programmed cell death (PCD), apoptosis is usually the first process that comes to mind. However, there is a new type of inflammatory PCD discovered in 2012, known as ferroptosis, that is genetically and biochemically distinct from other PCD.1
Anyone who is remotely interested in biology, or has perhaps scrolled through fitness websites to get in shape, has come across the word "protein". There is, however, much more to proteins than simply being a key player in maintaining active lifestyles. Proteins are ubiquitous in the cells of the body and are the driving force for key cellular processes. In order for proteins to carry out their duties, they need to be well-armed to execute their functions. This process of making the protein competent is achieved through specific post translational modifications (PTMs). The star of the PTMs is a cellular process called phosphorylation. The conventional methods adopted for quantifying phosphorylation are highly labor intensive. The development of phospho-specific antibodies has allowed for a huge sigh of relief from researchers due to their reputation of being quick, and detecting only phosphorylated forms of proteins in a complex mixture of phosphorylated and non-phosphorylated forms.
Oct 27, 2020 4:09:38 PM by Bryent Lee
A Closer Look at the Fundamentals of Recombinant DNA Technology
The advancement of recombinant DNA technology in recent years has drastically changed the world of research by controlling the expressions of target genes. Recombinant DNA combines genetic material from different sources, creating sequences that are unique and new to the genome. The DNA sequences used in the construction of recombinant DNA molecules can originate from any species, such as human, fungal, bacterial, and plants. 1
This has been an incredibly exciting past few days in the world of scientific research. For those unfamiliar, the 2020 Nobel Prize winners are set to be announced this week; as of today’s writing on October 7th, 2020, three sets of award winners have already been unveiled in medicine, physics, and chemistry. Today, we’ll take a closer look at the backgrounds and distinguished research of the laureates in medicine.
In a press release on August 26th, 2020, Abbott Laboratories announced that they were issued an emergency use authorization (EUA) by the FDA for their new, rapid point-of-care COVID-19 antigen test. Branded as the “BinaxNOW COVID-19 Ag Card”, the test is unique compared to the more prevalent molecular-based detection tests for COVID-19 in that it utilizes a lateral flow assay, similar to traditional over-the-counter pregnancy tests. Contained in a portable, credit-card sized device, Abbott claims that the test can deliver results in as little as 15 minutes, representing a significant reduction in time compared to RT-PCR-based tests that have turnaround times on a scale of hours rather than minutes.
For the first installment of our ABclonal Webinar Series on August 26th, we had the privilege of inviting Dr. Yong Xu of the Baylor College of Medicine to share his research on neuroendocrine mechanisms for appetite regulation and their implications on conditions such as obesity. If you were unable to attend the live session or would like to re-watch to review some of Dr. Xu's key points, we've got you covered with a link to a recording of the webinar and Q&A here, as well as recap of his lecture below.
Aug 19, 2020 7:50:57 PM by Dennis Miao
5 Ways to Stay Productive While Waiting for Experiments to Run
With labs across the country in various stages of reopening, it can be tough to transition right back into the physical office after months of working from home. Throughout your typical day in the lab, you may find that there are many instances where you've got a few odd minutes (or even hours) here and there. Oftentimes, these periods are used to scroll through your phone or catch up on social media. Here are five suggestions on how to productively fill in those gaps while you're waiting for your experiments to run.
When it comes to running a successful ELISA, there can be many common issues that must be addressed in order to obtain meaningful results. From our years of experience in producing and supplying ELISAs for a diverse range of targets, we’ve narrowed down four of the most common issues that can arise during your ELISA and have provided troubleshooting tips for each.
Whether it's at your home office, at the bench, or during your commute to work, putting on a podcast in the background can be an engaging and enriching way to help keep your mind stimulated. Here are 5 excellent science podcasts to help you get started on crafting your perfect workday playlist.
During these past few months, the push to discover effective treatments and vaccines against COVID-19 has largely overshadowed another vitally important aspect of drug development: drug pricing. Let’s take a look at a recent example from pharma giant Gilead, who recently announced the pricing for its promising COVID-19 treatment remdesivir.
In direct support of the fight against the novel coronavirus and the world's effort to understand more about the mechanisms of the COVID-19 disease, we're proud to announce the arrival of our SARS-CoV-2 Neutralizing Antibody.
Over these past few months, we here at ABclonal have been working diligently to continue supplying necessary reagents to researchers, clinicians, and companies worldwide to facilitate their efforts during the COVID-19 pandemic. In direct support of that goal, we’re proud to introduce our new, comprehensive line of SARS-CoV-2 ELISA kits.
On a bit of a lighter note this week, let’s take a dive into the origins of the beloved Guinea pig and their often overlooked role in the development of some major breakthroughs in the field of medical research.
As we’ve seen over these past few months, a SARS-CoV-2 infection can result in widely different manifestations and severities in the subsequent course of the disease it causes, COVID-19. Many of those infected by SARS-CoV-2 experience a mild to severe illness, with symptoms that include fever, shortness of breath, cough, and fatigue that appear roughly 2-14 days after exposure to the virus. On the other hand, some individuals infected with the virus will remain asymptomatic.
The novel coronavirus has spread rapidly around the world, with confirmed infection cases having reached more than six million. The average mortality rate of those infected is estimate to be around 6%. The sudden outbreak and particularly rapid spread of the novel coronavirus have required researchers worldwide to double down on investigative efforts against COVID-19 in order to develop a treatment and vaccine as soon as possible.
With troves of information and developments coming out daily from the various companies and researchers developing vaccines and potential treatments for COVID-19, it can certainly feel overwhelming to keep track of everything. This compilation of information should help you catch up on, and navigate recent developments in the fight against COVID-19.
As you may have surmised from the title of this article, Proteinase K (also known as protease K or endopeptidase K) shares many functional similarities to the protagonist of the iconic TV show, Breaking Bad. Much like Walter White, Proteinase K is incredibly versatile in its applications, while remaining relatively unassuming and overlooked at times. Unlike the chemistry teacher gone rogue, however, its properties can be channeled for good.
In recent days, there have been developments and progress towards initial re-opening of states across the country. For many of us, however, returning to work will be a more gradual process and will still involve some time at home as companies, localities, and the country as a whole work towards resuming normal life while maintaining social distancing and mitigating chances for secondary waves of infection. Thus, it’s still incredibly important to keep in mind these 5 strategies for working effectively, and healthily, at home:
May 7, 2020 6:20:55 PM by Dennis Miao
COVID-19 Detection Methods: Nucleic Acid vs. IgM/IgG Antibody Tests
There is no doubt that the coronavirus is the hottest topic as of late, having dominated media headlines and having fundamentally changed the way that we live and work. As the outbreak of the coronavirus continues to worsen across the globe, the demand for COVID-19 detection is therefore ever-increasing.
ABclonal has developed over 12,000 high quality antibody products since its inception in 2011. With this significant and time-tested experience, you can rest assured that ABclonal is relentless in its focus on the production and development of quality antibodies. Let’s further our understanding of antibodies by taking a more in-depth look at their structure, function, and uses in research.
The current COVID-19 public health crisis is unprecedented in the U.S. and worldwide. Everyone, including the U.S. Food and Drug Administration, medical workers, and biology researchers are busy doing whatever they can to fight against this epidemic. To help them handle the emergency and buy time for the suffering patients, ABclonal developed ABScript II One Step RT-qPCR Probe Kit (RK20407), a ready-to-use kit that can be used to quantify RNA with outstanding performance. Here, we would like to introduce the kit as well as the four reasons that you should use it.
The current coronavirus (COVID-19) pandemic is an unprecedented public health emergency in the U.S. Even though medical professionals are working around the clock to conduct testing, the problem still remains to be a shortage of diagnostic kits. Aimed to improve the limited diagnostic capability and to fight against the COVID-19 public health crisis, the FDA issued several emergency policies since late February.
Do you find yourself using a lot of SDS-PAGE gels for Western blotting and Coomassie staining? Or have your pre-cast gels been stored for too long and expired? Why not try to cast your own SDS-PAGE gels to save some budget for the lab, and produce just as valid of results. Today, we would like to share five tips for hand-casting SDS-PAGE gels, as well as the protocol and formulation to do so.
Autophagy is a natural mechanism in which the cell removes and degrades cellular components with autolysosomes. It is a popular research area because autophagy is related to many physical and pathological processes. The 2016 Nobel Prize in Physiology is granted to Yoshinori Ohsumi for his contribution in autophagy. In autophagy studies, LC3-I and LC3-II detection is a must-have experiment to track the autophagy process. Therefore, we would like to share five important notes for quantifying autophagy with LC3.
The Wnt signaling pathway, an evolutionarily conserved signal transduction pathway, is widely present in invertebrates and vertebrates. The Wnt signaling pathway plays a crucial role in early embryonic development, organogenesis, tissue repair, and many other physiological processes. The mutation of key proteins involved in this pathway can lead to abnormal activation of signals, and potentially induces the occurrence of cancer. In 1982, R. Nusse and H.E. Varmus identified the first Wnt gene from a mouse mammary tumor and named it Int1 (integration 1). Continued research found that the mouse Int and Drosophila Wingless (Wg) genes are homeotic, and thus combined their names to Wnt. H.E. Varmus himself also won the 1989 Nobel Prize in Physiology or Medicine for his great contribution in oncology.
Western blot results can either be the highlight of the day, or a scientist's worst nightmare. Unfortunately, it usually turns out to be the latter of the two. After a long hard day of work, nothing can ruin the day more than seeing your western blot results come out blotchy and unreadable. Luckily, there are many ways to prevent and fix a messy blot to ensure you get the best results possible, rather than ending up with something horrific like the image below. To avoid these situations, I have outlined some tips to keep in mind before going through with your western blot test.
Oct 31, 2019 3:31:35 PM by Daniel Bouzas
2019’s Top 5 Breakthroughs in Breast Cancer Research & Treatment
Every year about 12% (one in eight) women across the U.S. will be diagnosed with breast cancer, with an estimated 41,760 expected to succumb to this deadly disease by the end of this year alone. However, new milestones in research is helping us understand more about how we can further improve treatment for those currently battling through breast cancer in hopes of increasing the overall survival rate. Thankfully, every year since 2000 we have seen a 7% decrease in reported incidents, and in 2019, milestones in how we treat different types of breast cancer show hope for this number to increase in the future. With breast cancer awareness month coming to an end, we wanted to shine a light on this year's biggest developments in research and available treatments.
Have you ever felt unprepared for a job interview or presentation? Everyone has at one point, but while some might crack under pressure due to that nervous feeling, others will go through with it with no sweat. How do they do it? By keeping an elevator pitch in mind as a guide for what to say and when to say it.
We want you to go in and out of every meeting feeling confident in your performance, so below are some crucial tips to keep in mind when developing an elevator pitch of your own.
Humans have been into music ever since the first group of cavemen started banging sticks on rocks and stone walls, so I guess you could say we got music engraved in our genes. However, as researchers would find out later, there could be some hint of truth to that statement. Thanks to advances in the way we read genetic coding, your DNA can be converted into listenable frequencies.
Sep 24, 2019 1:45:02 PM by Daniel Bouzas
The Struggling Bio-Graduate Guide To Finding The Right Path
When you’re just starting or nearing the end of your college career, you are either feeling prepared for the adult world ahead, or you still feel lost about what are the next steps in your professional career. If you’re reading this, I’m guessing you might fall in the latter, and I’m sure you might have questions regarding the many paths you can take in this biotech industry. For some jobs, the road to success is a clear one, but for the biotech industry, it could be a lot harder to decide what direction to take due to the many options that are laid out for you. For this blog post, I wanted to provide a one-stop guide answering FAQs that I gathered from graduates going through the same struggles as you are.
It’s a relatively new world for scientists. Up until the 2000s, research funding increased steadily before reaching a plateau and dropping with sequestration budget cuts. Nowadays, scientists spend a great deal of time fighting for grants, rather than actually doing research. It’s an interesting, but sobering reality: as you progress in your science career, you may (or already do), find yourself spending more and more time planning and writing grants.
Cell proliferation assays have a wide range of applications in scientific research – from testing drug reagents to the effect of growth factors, from testing cytotoxicity to analyzing cell activity. So, what are cell proliferation assays? Cell proliferation assays typically detect changes in the number of cells in a division or changes in a cell population.
In a previous article, we explored the differences between rabbit and mouse antibodies as well as the biology behind rabbit antibody superiority. But after choosing the host, the type of technology used to produce the antibody is important too. Here, we explore some of the rabbit monoclonal antibody technologies available in the current market.
Aug 29, 2019 2:57:57 PM by Michele Mei
Reproducibility Crisis: Fallacies to Be Wary of and Ways to De-Bias
While the scientific community is enveloped in a reproducibility crisis (and debates as to whether there is one), there are certainly steps life science researchers can take to ensure more reproducible outcomes. We can start by limiting self-bias and improving reporting standards. But first, what is reproducibility and why is there a crisis?
Podcasts are perfect for busy, but intellectually curious people. It’s like reading, but instead of fixing your eyes to a page or screen, you can run or cook or simply relax while the podcast delivers fascinating, funny, new information straight to your brain. It’s basically like learning by osmosis! Whether you’re working in lab or you have these few months off to relax, I curated a list of science podcasts to keep you company both bench-side and poolside.
Large-scale sequencing projects have great potential to provide a wealth of knowledge to scientists and the public. Perhaps the most celebrated project of this nature is the Human Genome Project (HGP) which was completed in 2003. For many, the multi-billion endeavor was considered a “moonshot” for biology, but with its successful completion (99% of the euchromatic genome sequenced with 99.99% accuracy) came the launch of many other large-scale sequencing projects such as the Cancer Genome Atlas (2005) or more recently, the Earth Bio-genome Project (2018). The introduction of large-scale quantitative methods, such as next-generation sequencing, have also made these projects feasible.
As one of the most common reagents in biology and medical research, there are more than 350,000 commercially produced antibodies available for research and clinical applications. However, the quality of the commercially available antibodies varies from vendor to vendor. Different suppliers have different protocols for validating antibodies and some researchers might want to verify the product before using them on precious samples. Here are some of the factors to examine when it comes to antibody quality.
A healthy immune system requires a series of checkpoints to ensure self tolerance and prevent damage to other tissues during immune response. Binding of costimulatory signal transduction molecules (such as CD28, ICOS, GITR) on T cells to their receptors (such as CD80/CD86, ICOSL, GITRL) on antigen presenting cells (APCs) may contribute to T cell activation. However, in some states, inhibitory signals of T cell activation and response occur during the involvement of T cell receptors. These signals are generated by proteins involved in immune checkpoints (eg, PD-1, CTLA-4, TIM-3, and LAG3). Usually PD-1 and CTLA-4 immunological checkpoint proteins are upregulated in T cells infiltrating tumors and bind to their respective ligands, PD-L1 (ligand B7-H1)/PD-L2 (ligand B7- DC) and CD80/86, and down-regulate T cell responses. Immunological checkpoint ligands are often upregulated in cancer cells as a means of evading immune detection. Therefore, immunotherapy by blocking immunological checkpoint protein activation of anti-tumor immunity has become a popular research subject for cancer therapy.
The cliché of the pragmatic and lonely scientist gets old. Although scientists are highly analytical, their emotional range is not as limited as the media and stereotypes portray. In their work, scientists must be logical and methodical, but that doesn’t necessarily carry over to life and relationships.
The interest in using primary cells for cell-biology research has gained prominence in recent years due to factors such as cell line contamination (Kaur G, 2012). What made primary cells lose their popularity in the first place is partly due to the rigorous and arduous process associated with primary-cell cultivation. So why is primary-cell cultivation so difficult?
It’s no secret that scientific research is becoming less of a priority to the federal government. For two decades, research and development (R&D) funding has remained stagnant or dropping, despite increases to the overall federal budget. With a growing population of scientists entering the field, a lack of funding generates a hyper-competitive and stressful funding climate. For those looking to secure funding for the first time, or simply curious about how science is funded, this post serves as an introductory guide.
Being perpetually busy has become a status symbol in academia –and it’s counterproductive.
In this day and age, we are trained to believe that the more you work, the more you get done, and the further ahead you get. In academia, researchers place a lot of pressure on themselves to work around the clock. Whether it’s experiments, teaching, papers, or grants, it seems like there’s always more to be done. Consequently, the lack of work-life balance, work-induced stress, and burnout has become a pervasive problem in academia.
The G2/M cycle checkpoint prevents cells with genomic DNA damage from entering mitosis (M phase). The main safeguards conferred by this checkpoint is to ensure that DNA is free of major lesions or replication errors, and there are enough organelles, metabolites, and other cellular cargo in the parent cell prior to division so the daughter cells can be adequately provided for once mitosis is complete. Failures at this checkpoint are associated with aberrant cellular growth and cancer progression.
The Cyclin B-CDK1 complex plays an important regulatory role during the G2 transition, at which time CDK1 is maintained inactivated by the tyrosine kinases Wee1 and Myt1. When the cells enter the M phase, the kinase Aurora A and the cofactor Bora act together to activate PLK1, which in turn activates the activity of phosphatase CDC25 and downstream CDC2, effectively driving the cells into mitosis. When the DNA is damaged, it activates the DNA-PK/ATM/ATR kinase and eventually inactivates the Cyclin B-CDK1 complex. Stopping cell cycle progression allows the cell enough time to attempt to repair any DNA or cellular damage, and if all else fails, to induce apoptosis to prevent risk to the entire organism.
ABclonal Technology provides a wide selection of cell cycle checkpoint antibody products for every phase of a cell's life. Please see a small sample of our offerings below.
In some ways, the heart is quite a vulnerable organ. Cardiac complications such as heart attack, cardiac arrest, or heart failure are common. But interestingly, of the many diseases that may affect the heart, cancer is not one of them. For example, we often hear about cancer in the prostate, breast, colon, skin, etc., but rarely of the heart. How is this vital organ different?
The G1/S cell cycle checkpoints control whether eukaryotic cells enter the S phase (synthesis phase) of DNA synthesis after having properly completed the G1 phase to ensure the cell has enough energy and resources to begin DNA replication. Two cell cycle kinase complexes, CDK4/6-Cyclin D and CDK2- Cyclin E, work together to relieve the inhibition of dynamic transcriptional complexes containing retinoblastoma protein (Rb) and E2F. In cells undefined during the G1 phase, hypophosphorylated Rb binds to the E2F-DP1 transcription factor and forms an inhibitory complex with HDAC, thereby inhibiting downstream key transcriptional activities. Clear entry into the S phase is achieved by continuous phosphorylation of Rb by Cyclin D-CDK4/6 and Cyclin E-CDK2, which separates the transcription factor E2F from the inhibitory complex and allows transcription of the gene required for DNA replication. After the growth factor disappears, the expression level of cyclin D is down-regulated by down-regulation of protein expression and phosphorylation-dependent degradation. Without a proper G1/S checkpoint, the cell could arrest or potentially undergo aberrant processes that could lead to disease states such as cancer.
Proteins known as transcription factors play a crucial role in gene regulation by activating, enhancing, and even silencing a gene’s expression. Many textbooks and resources compare transcription factors (TFs) to something like an on/off switch for gene transcription. However, it is a bit more complicated than just turning gene expression on or off. Various properties (e.g. binding affinity, specificity, and genetic variance of binding sites) impact the binding of TFs to DNA, thereby altering gene expression. To study transcription and how it is regulated, scientists study TF-DNA interactions on a genome-wide level.
Embryonic stem cells (ES cells) are pluripotent stem cells isolated from an inner cell mass of early-stage embryo-blastocysts. ES cells have a high differentiation potential., which means that they have the capacity to develop into whatever cell type the body needs depending on the signals received by the ES cell. At the same time, while ES cells are undifferentiated, they retain the potential to infinitely replicate, making them highly attractive and renewable subjects for targeted cell therapy and regenerative medicine.
Cluster of differentiation, or CD molecules, are cell surface markers that are used for identification of cell types in pathology and other bioscience disciplines. The expression levels of CD markers may increase or decrease (or disappear altogether, at least to undetectable levels) when cells (for example, leukocytes, red blood cells, platelets, and vascular endothelial cells, etc.) differentiate into new and different lineages. Depending on the CD marker, the expression level may identify a phenotype for different segments of cells, such as when they become active or diseased. Most CD molecules are transmembrane proteins or glycoproteins, including extracellular regions that bind a ligand or opposing receptor, transmembrane regions to anchor the CD marker into the cell, and cytoplasmic regions that may confer some adaptor or catalytic function. Some CD molecules can also be "anchored" on the cell membrane by means of inositol phospholipids. A few CD molecules are carbohydrate haptens. The study of CD molecules can be used in many basic immunology research fields, such as the relationship between CD antigen structure and function, cell activation pathway, signal transduction and cell differentiation, etc. It can be used clinically for disease mechanism research, clinical diagnosis, disease prognosis, efficacy tracking and treatment, and more. CD molecules such as CD4, CD8, CD25, etc. can be used to identify populations of cells when studying samples by flow cytometry or immunofluorescence.
The Literature Review
Literature reviews are some of the most widely read and highly cited papers in academia, but writing one can be a daunting task, requiring an expert understanding of the topic at hand. To write a review article is so much more than simply summarizing recent studies published in the field. The most valuable literature reviews, which I find myself going back to again and again, are those that:
The Problem with Cancer Models
Very few cancer drugs succeed in clinical trials, despite showing promise in the lab. Treatments that may work on animal models, cell lines, or even patient-derived xenografts often do not have the same efficacy in patients. The underlying reason is tumor environments within the human body are far more complex than in research models. For example, the tissue structure (histological complexity) and genetic heterogeneity of an animal model is different than that of humans. Even cell lines and patient-derived xenografts, which are human-derived, have their own pitfalls such as genetic mutations and animal-specific tumor evolution, respectively. Due to the inability to reproduce human tumor environments, many drugs fail clinical trials after lengthy and costly development.
The Hippo signal is very conservative in evolution. It regulates organ size and tissue stability by regulating cell proliferation, apoptosis, and stem cell renewal. The core process of Hippo signaling is a kinase tandem process, Mst1/2 and Sav1 form a complex, phosphorylate and activate Lats1/2; Lats1/2 kinase then phosphorylates and inhibits transcriptional coactivators Yap and Taz. Yap and Taz are the most important effectors downstream of the Hippo pathway. Upon dephosphorylation, Yap and Taz translocate to the nucleus and interact with TEAD1-4 or other transcription factors (such as CTGF) to induce gene expression, thereby initiating cell proliferation and inhibiting apoptosis.
As scientists, writing is a major component of the job, yet having “no time to write” is a common complaint echoed amongst PhD candidates, post-docs, and professors alike. On top of experiments, data analyses, and taking/teaching courses, writing can easily end up on the back burner. But publishing papers, like it or not, is critical for a career in science. Rather than setting intimidating goals like publishing some number of papers within a year or publishing in a high impact journal, it is more feasible and beneficial to first develop good writing habits, which will in the long run increase productivity.
Every year, scientists make fascinating breakthroughs which broaden, yet challenge, our understanding of life and the world around us. Just as we start to understand a biological process, like how heredity or aging works, a new discovery can flip it on its head or open a whole new avenue for research. As 2018 comes to an end, it’s the time for roundups of top products, gifts, movies, tech, etc. We decided to put our own spin on it with the top life science discoveries of the year.
These days major debates center around scientific information – from climate change, gene-editing to vaccinations – yet, despite the data-driven nature of science, there are deeply divided opinions regarding these hot topics. For researchers, it might be frustrating to witness scientific findings being misinterpreted or exaggerated. But it’s not surprising that so much science is misunderstood. Too many scientists still reside within their own research bubbles, which is counterproductive.
The glyceraldehyde-3-phosphate dehydrogenase, or GAPDH for short), is a multifunctional, indispensable enzyme found in all cells. The generally known function of GAPDH is to assist in carbohydrate metabolism as a key player in glycolysis, but there are studies demonstrating its role in the nucleus as well.
GAPDH is a constitutively expressed housekeeping protein, and GAPDH mRNA levels and protein levels are often used as loading controls in experiments that quantify target-specific expression changes. Recent studies have elucidated the role of GAPDH in apoptosis, gene expression through its possible activities as a transcription factor, and nuclear transport. As both a metabolic protein as well as one that might play a role in cytoskeletal reorganization, GAPDH activity is intricately tied to tumorigenesis. GAPDH may also play a role in neurodegenerative diseases such as Huntington's disease and Alzheimer's disease. Therefore, although many researchers do use GAPDH as a control, this protein needs to be appreciated for its myriad other functions as well!
ABclonal Technology's GAPDH recombinant rabbit monoclonal antibody is a human-specific antibody that can be used with a high dilution ratio of 1:2560000. As a highly-stable antibody product, this means that you can perform numerous Western blotting experiments over a long period of time using a small quantity of antibody, as well as in other experiments to study the functions of GAPDH. Take advantage of this robust, cost-effective antibody product in your research today!
In the last What’s Hot in Life blog post, we discussed how next generation sequencing (NGS) is used as a basis for understanding disease. This week I wanted to talk about DNA sequencing again, but in a completely different context. On November 1st, scientists launched an ambitious project to sequence all 1.5 million complex species on Earth. Their purpose? To save biodiversity.
Nov 28, 2018 4:22:23 PM by Panyue (Penny) Hao
Scientists Identify Novel Regulator for LINE-1 Using ABclonal Antibody
Long-interspersed nuclear elements (LINEs) are genetic components found in higher eukaryotes. They are retrotranposons, meaning that they are transcribed into mRNA and then translated into proteins that act as a reverse transcriptase. The reverse transcriptase makes a copy of the LINE DNA which can then be integrated into the genome at a new site. The only active LINE in humans is LINE-1. It has been associated with oncogenesis and Haemophilia A, a diseased caused by insertional mutagenesis.
Although underappreciated, the Golgi apparatus is indispensable to normal cellular function by ensuring proteins are properly folded and sorted, and to direct diverse functions including autophagy. Disruptions to proper Golgi function can lead to many disease states, including diabetes, cancer, and neurodegenerative disorders such as Alzheimer's disease. The study of vesicular markers, including Golgi markers, is critical to our understanding of this amazing organelle's function in keeping the cell and organism healthy. Clarifying the mechanisms by which proteins are properly folded, sorted, quality controlled, and transported will prove important as more effective therapies are developed against a diverse array of human diseases.
We have previously explored the function of organelle markers USO1, GOLGA2, and GOLM1 but not how the corresponding antibodies can be applied in research. Organelle marker antibodies are common tools in cell biology research. They can be used with immunofluorescence technology to observe the morphological structure of organelles and understanding the subcellular localization of proteins. In turn, they help to explore the biological functions/role of organelle proteins in normal or disease models. These markers can also be used in Western blot (WB) experiments examining organelle extracts: as a positive control to determine whether the organelle is successfully extracted.
You can see some examples of ABclonal Technology's Golgi marker antibodies below. These are only a handful of the huge selection of targets that you can use to supplement your cutting-edge research!
Pursuing a PhD is undoubtedly one of the most challenging chapters in a researcher's career. For the first time, as an early career scientist, you must juggle research, writing, teaching, and your own personal life (yes, you should still have one). A PhD is definitely exhausting, but given the right guidance and support it can be an enjoyable and exciting time too.
Organelle marker antibodies are common tools in cell biology research. They can be used with immunofluorescence technology to observe the morphological structure of intracellular membrane-bound organelles and for understanding the subcellular localization of proteins. In turn, they help to explore the biological functions/role of organelle proteins in normal or disease models. These markers can also be used in Western blot (WB) experiments examining organelle extracts, as well as providing a positive control to determine whether the organelle is successfully extracted.
We focus today on the endoplasmic reticulum (ER) markers. The ER is a network of membrane-bound organelles that are the initial destination for proteins that are targeted for other organelles, the plasma membrane, or are to be secreted outside the cell. Proper ER function includes accepting the nascent protein as it is being translated by ribosomes, and ensuring that the protein is properly folded so it does not lead to accumulation of unusable cellular cargo. Disruptions in ER or the unfolded protein response can lead to cell death or various disease states such as cancer or neurodegenerative disorders.
ABclonal provides many antibody products for the study of ER and Golgi markers, as well as exosome markers. Please read our blog on the vesicular transport system and its role in cellular homeostasis, and check out some of our ER marker antibodies below. We are honored to be part of your journey to better understanding vesicular transport and the fight against human diseases!
Oct 30, 2018 5:56:54 PM by Dapeng Sun, Ph.D.
Choosing the Right RNA Lib Prep Kit for Your Transcriptome Library
1. Determine the mRNA capture type you need
The scientific community operates on a self-correcting model that relies on repetition and replication. However, according to a 2016 survey by Nature, more than 70% reported to have failed to replicate experiments from another scientist, more than 50% reported failure in replicating his/her own experiment. Out of the 1,576 scientists surveyed, 906 were from biology or medicine disciplines.
CTCF (CCCTC Binding Factor) is a highly conserved transcription factor that regulates transcriptional activation, transcriptional repression, insulator function, and imprinted control regions (ICRs).
Therapies targeting the function of a small intestinal protein, SGLT1, might have the potential to treat diseases like obesity, diabetes, heart failure, and associated death—and we have next generation sequencing to thank.
CREB1 is a basic leucine zipper domain (bZIP) transcription factor that activates a target gene through a cAMP response element. As a key transcriptional regulator, CREB1 plays a role in a variety of cellular responses by mediating a number of physiological stimuli. CREB1 is expressed in many tissues and plays an especially important regulatory role in the nervous system by promoting neuronal survival, driving precursor proliferation, neurite outgrowth, neuronal differentiation and more. In addition, CREB1 signaling is involved in the learning and memory functions of many organisms. CREB1 is capable of selectively activating many downstream genes through interaction with multiple dimerization partners. Phosphorylation of CREB1 at the serine 133 site involves multiple signaling pathways, such as Erk, calcium flux (Ca2+), and stress signaling. Some of the kinases involved in CREB1 phosphorylation include p90RSK, MSK, CaMKIV, and MAPKAPK-2.
Announcements for this year’s Nobel winners started off with prizes in Physics, Chemistry, and Physiology or Medicine. While congratulations are in order for the newly minted laureates, a bit of controversy is also stirring.
Autophagy is a catabolic process in which autophagic lysosomes, known as autophagosomes, degrade most cytoplasmic contents, including entire organelles like damaged mitochondria in protection of the host cell and organism. Autophagy is usually activated in the absence of nutrients and is associated with many physiological and pathological processes, including growth, differentiation, neurodegenerative diseases, infections and tumors. Light chain 3 (LC3) is a widely recognized autophagy marker. There are three isoforms of the LC3 protein (LC3A, LC3B, and LC3C) in mammals. They undergo post-translational modifications during autophagy. The LC3 protein is first cleaved by Atg4 at its carboxy terminus immediately after synthesis to produce LC3-I, which is localized in the cytoplasm. During autophagy, LC3-I is modified and processed by a ubiquitin-like system including Atg7 and Atg3 to produce LC3-II with a molecular weight of 14 kD and localized to autophagosomes. The magnitude of the LC3-II/I ratio can be used to assess the level of autophagy.
ABclonal Technology is pleased to offer numerous antibody products to study targets within the autophagy pathway, be it the normal functions within the pathway, the autophagic cell death pathway, or disruptions to autophagy that may drive disease progression. Many of our products have been peer reviewed by satisfied customers, including those who have used them to generate quality data for scientific publications. Please see some examples of our products below, and happy experimenting!
ABclonal Technology hosted its second lunch and learn at the Koch Institute for Integrative Cancer Research at MIT, the second event of its lecture series. The lunch and learn, led by ABclonal’s senior principal scientist, focused on rabbit monoclonal antibody technologies, its advantages and development.
Sep 25, 2018 12:20:19 PM by Panyue (Penny) Hao
RNA Methyltransferase Antibody Featured in Cell and Nature Journals
RNA methyltransferases such as METTL3, METTL14, WTAP, and VIR can catalyze the methylation of the N6 position of adenylate (M6A) and are opposed by demethylases which include FTO and ALKBH5.
The nucleosome consists of an octamer composed of four histones (H2A, H2B, H3, and H4) and a DNA entangled with 147 base pairs. The core of the histones constituting the nucleosome are roughly the same, but the free N-terminus can be subjected to various modifications.
Sep 11, 2018 10:58:30 PM by Panyue (Penny) Hao
Featured Product Weekly: ER and Nuclear Membrane Markers
Endoplasmic Reticulum Marker
The endoplasmic reticulum is a membrane-bound organelle that is critical to the proper sorting and folding of proteins. Improperly folded proteins are normally allowed to refold into their functional conformation, and if not possible to repair, these unfolded proteins are directed to be degraded to prevent damage to the cell. The P4HB gene encodes a protein disulfide isomerase (PDI) that catalyzes both the formation of disulfide bonds, which form between cysteine residues to stabilize protein structure, and isomerization between or within molecules of secreted proteins. To achieve the natural conformation, this process takes place in the endoplasmic reticulum, so P4HB is often used as an ER marker. Studies on the oxidative folding mechanism indicate that molecular oxygen can oxidize the ER protein Ero1, and Ero1 can oxidize PDI through a disulfide bond. After this activity, PDI catalyzes the folding of proteins to form disulfide bonds.
Ebola outbreaks are considered rare, but they do emerge every several years and can be quite lethal. Although the first confirmed Ebola epidemic was in 1976, we still lack licensed therapeutics to prevent and control Ebola’s spread. Vaccine development is in the works, but the lack of an approved treatment is a chilling reminder that we may not know enough about the virus. With the recent outbreaks in mind, we sought to summarize everything you should know about Ebola, its biology, and the current progress of vaccine development.
The extracellular signal-regulated kinases, or ERK1/2 (MAPK1/MAPK3, p44/42MAPK), are signaling molecules belonging to the mitogen-activated protein kinase family (MAPKs) that are commonly located in the cytoplasm of eukaryotic cells. In concert with various other molecules in the signaling cascade acting under different surface or intracellular receptors, ERK1/2 act as catalysts in the phosphorylation of serine/threonine and are negatively regulated by the bispecific (Thr/Tyr) MAPK phosphatase family (called DUSP or MKP) and specific inhibitors to MEK activity (such as U0126 and PD98059).
When I began my science journey as an undergrad, research seemed rigorous, but reassuringly straightforward in its tenets. Observe, question, hypothesize. Predict, test, analyze. And repeat. It made perfect sense to me that if you followed this protocol and remained unbiased in the process, great discoveries were sure to come.
But then I learned about the other steps in between. Steps like grant-writing, worrying about publishing and impact factors, getting your mentor to actually respond, and struggling to troubleshoot experiments. Twitter’s PhD community seems to relate.
Aug 28, 2018 9:00:00 AM by Panyue (Penny) Hao
Featured Product Weekly: DNA Methyltransferase Antibody
DNA methyltransferase (DNMT) is an important family of enzymes that catalyze and maintain DNA methylation, a common marker in the epigenetic silencing of target genes. The enzymes play a key role in the regulation of gene expression and genomic imprinting/development.
Astrocytes are specialized glial cells with distinct morphology that are found in the central nervous system, playing a role in brain and nerve cell development and the formation of synapses. Mature astrocytes respond to many stress signals and are responsible for many essential complex functions in the healthy brain, allowing the maintenance of proper homeostasis through ion flow, signaling, and the recycling of neurotransmitters. Astrocytes that are irregularly activated may result in various neurological disorders, including Alzheimer's disease and Huntington's disease.
Glial fibrillary acidic protein (GFAP) is an intermediate filament protein that is mainly found in astrocytes. It is also expressed in chondrocytes, fibroblasts, myoepithelial cells, lymphocytes, and hepatic stellate cells.
Epidermal growth factor receptor (EGFR, also known as ErbB-1 or HER1) is a member of the ErbB family. This family includes four tyrosine receptor kinases: HER1 (ErbB1, EGFR), HER2 (ErbB2, NEU), HER3 (ErbB3), and HER4 (ErbB4). The ErbB family plays an important regulatory role in the process of cell physiology, and is among the most studied receptor tyrosine kinases and signaling molecules in the history of biochemistry and cell biology.
EGFR is distributed along the surface of cells including mammalian epithelial cells, fibroblasts, glial cells, keratinocytes, and more. The EGFR signaling pathway plays an important role in physiological processes such as cell growth, proliferation and differentiation. Upon ligand binding (for example, EGF interacting with the extracellular domain of EGFR), the ErbB receptor tyrosine kinases will homodimerize or heterodimerize, allowing autophosphorylation of cytosolic tyrosine residues and the recruitment of downstream signaling molecules.
The loss of function in tyrosine kinases such as EGFR, or the abnormal activity/cell localization of key factors in related signaling pathways, such as the p38 MAPK pathway, can cause multiple cancer types, diabetes, immunodeficiency and cardiovascular diseases. In modern medicine, typical treatment strategies include targeting the tyrosine kinase activity of the ErbB receptor with small molecular inhibitors, or humanized antibodies that will target cells that have overexpressed the ErbB receptor, such as using an anti-HER2 therapy to treat breast cancer.
Although underappreciated, the Golgi apparatus is indispensable to normal cellular function by ensuring proteins are properly folded and sorted, and to direct diverse functions including autophagy. Disruptions to proper Golgi function can lead to many disease states, including diabetes, cancer, and neurodegenerative disorders such as Alzheimer's disease.
Part of the Golgi protein family, USO1 protein (also known as vesicle docking protein p115) is a peripheral membrane protein that can be used as a Golgi marker. It cycles between the cytoplasm and the Golgi apparatus during interphase. The position of the USO1 protein is regulated by phosphorylation -- dephosphorylated proteins bind to the Golgi membrane and dissociate from the membrane when phosphorylated. This regulated transportation plays an important role in protein localization, secretion, and signal transduction. USO1 protein acts as a vesicle anchor by interacting with the target membrane and keeping the vesicles close to the target membrane. In addition, the USO1 protein interacts with GOLGA2 (GM130) and Giantin to promote endoplasmic reticulum-Golgi transportation. A large part of Golgi-related research is in understanding how to maintain proper Golgi function to prevent and treat human diseases.
AIFM1, also known as Apoptosis Inducing Factor (AIF), is a widely expressed flavoprotein that plays an important role in caspase-independent apoptosis. AIF normally exists in the mitochondrial intermembrane space.
Nuclear lamina is a layer of cross-linked fibrin network that commonly exists in higher eukaryotic cells. It is interior to the nuclear envelope with a fiber diameter of about 10 nm. The nuclear lamina of higher animals are usually composed of three intermediate filament polypeptides – lamins A, B, and C. The nuclear lamina is closely related to the stability of nuclear envelopes, maintenance of nuclear pore location, stabilizing interphase chromatin morphology and spatial structure, chromatin construction, and nuclear assembly.
ABclonal Technology began its lecture series at Sanofi Pasteur, located in Cambridge, Massachusetts. The lunch and learn event focused on rabbit monoclonal antibodies in the current market, which is one of ABclonal's leading product lines to support biological research.
Golgi membrane protein 1 (GOLM1) is a type II Golgi membrane protein discovered in recent years. Although underappreciated, the Golgi apparatus is indispensable to normal cellular function by ensuring proteins are properly folded and sorted, and to direct diverse functions including autophagy. Disruptions to proper Golgi function can lead to many disease states, including diabetes, cancer, and neurodegenerative disorders such as Alzheimer's disease. GOLM1 is a key protein in ensuring that proteins taken in from the endoplasmic reticulum are properly transported to their final destination in and out of the cell, and also may be involved in responses to viral infection.
The GOLM1 protein expression level increases in a variety of diseases and cancerous tissues. It is especially closely related to liver diseases. Many studies have shown that GOLM1 is more sensitive and more specific than alpha-fetoprotein (AFP, the most specific marker for primary liver cancer and the main indicator for the diagnosis of liver cancer) in the serological diagnosis of liver cancer. GOLM1 is expected to be the serological marker for the early diagnosis of liver cancer. It has also been reported to be highly expressed in patients with viral hepatitis and cirrhosis. The study of GOLM1 and other Golgi markers remains crucial to our understanding of human disease, and offers new avenues to develop more effective targeted therapies to alleviate the burdens of these ailments.
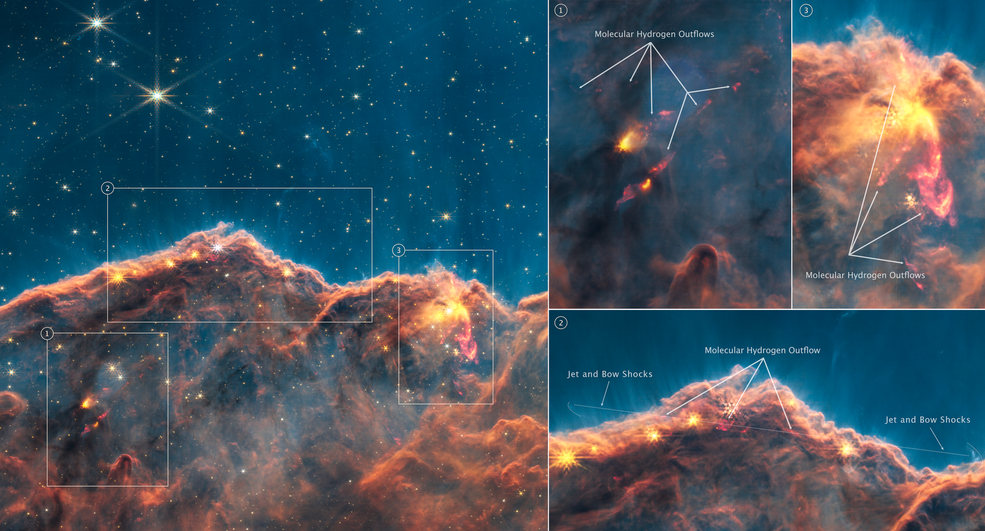





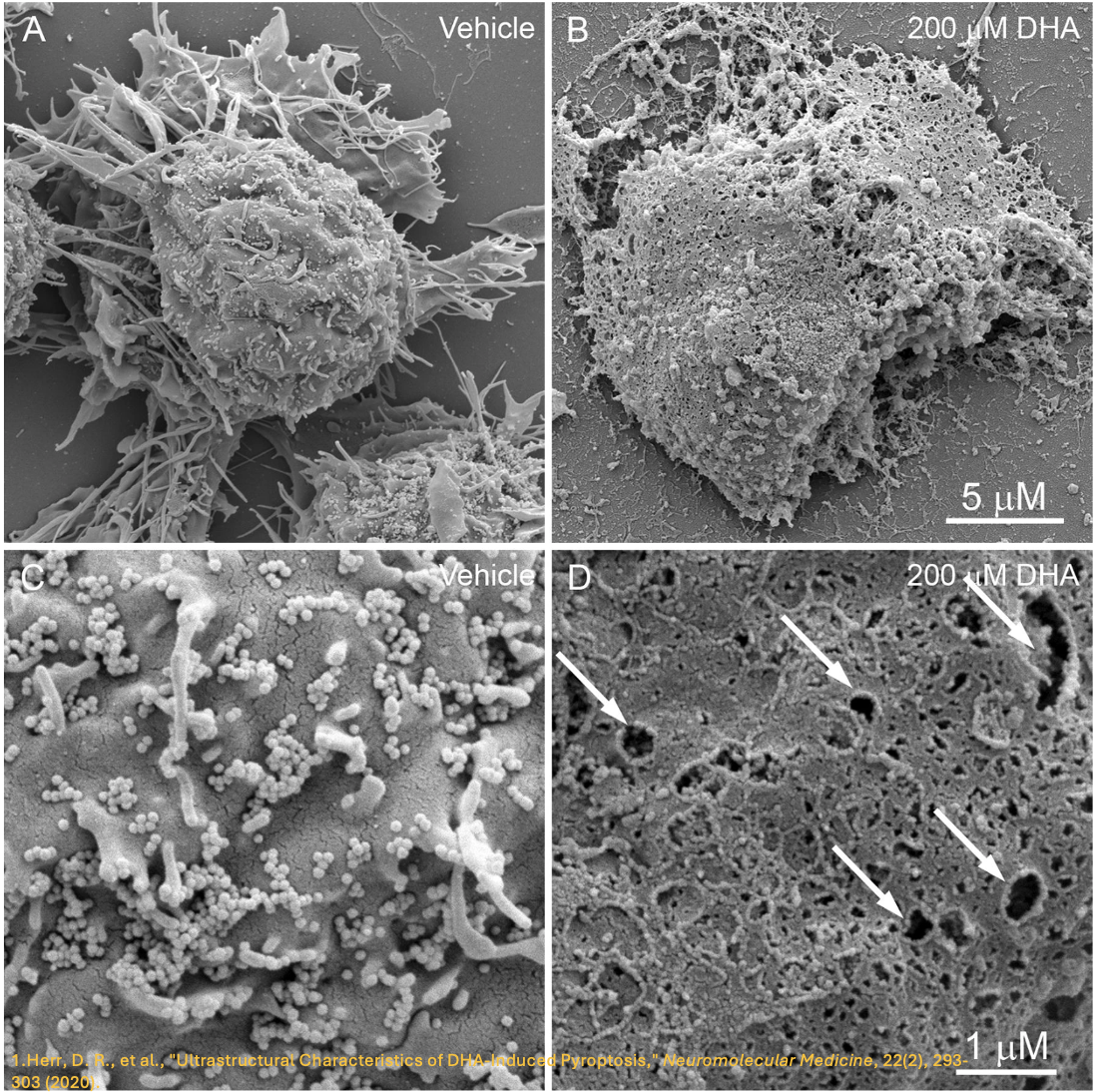

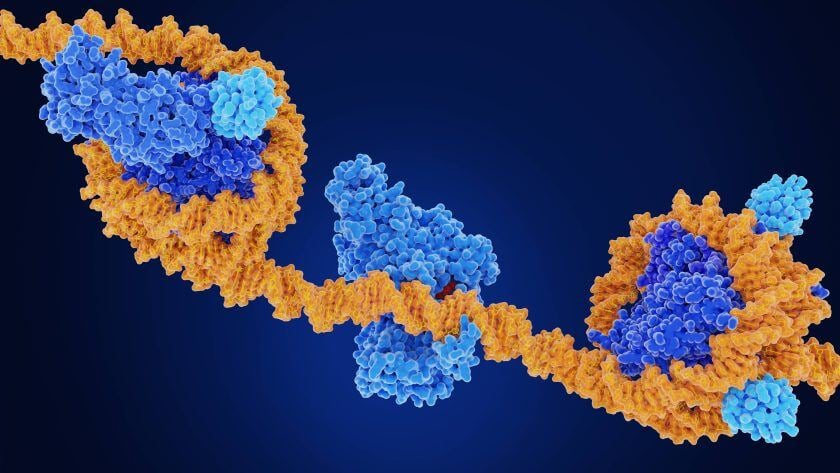







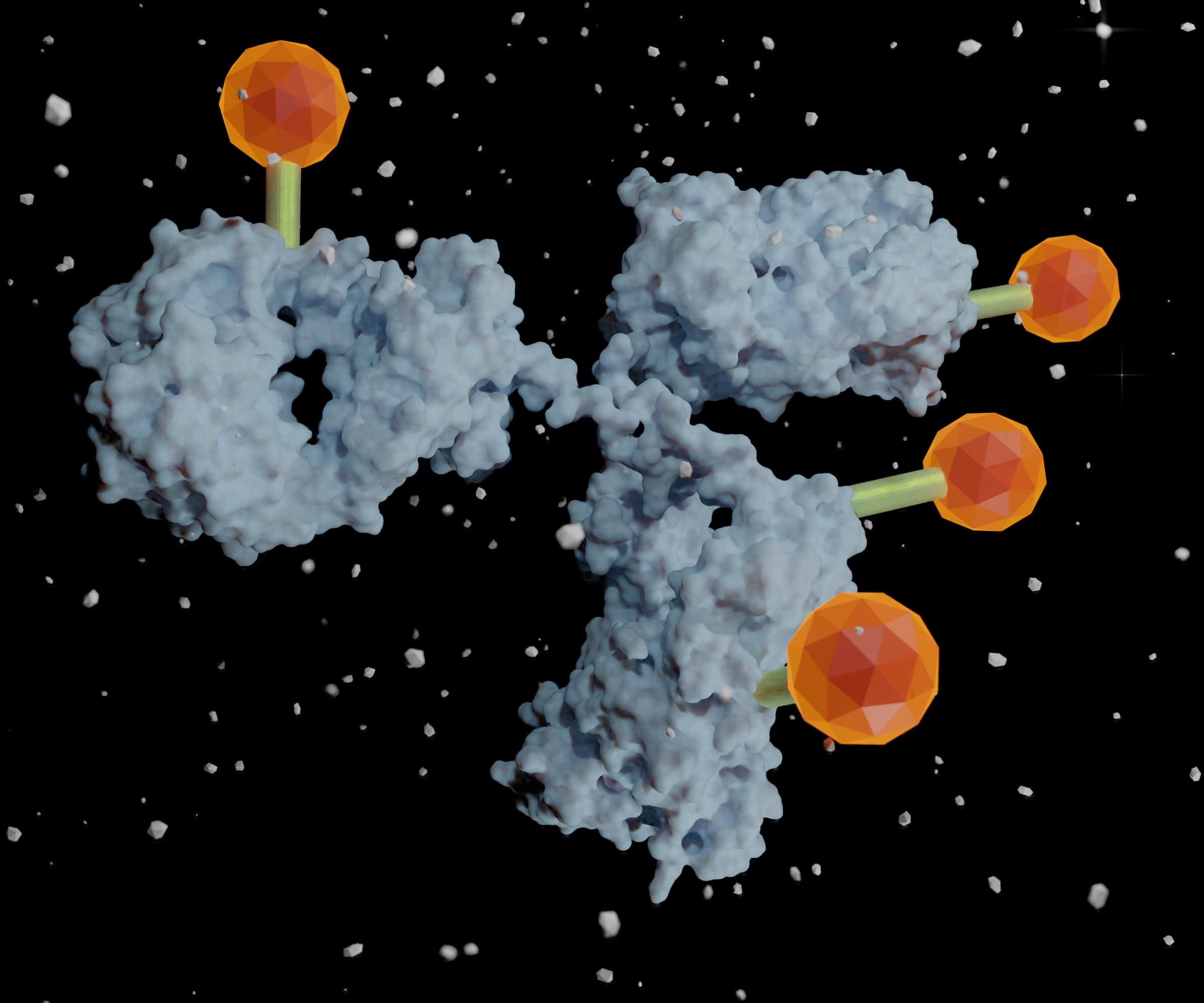







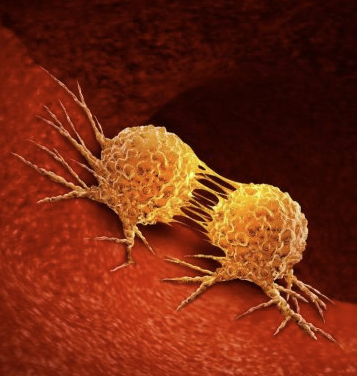



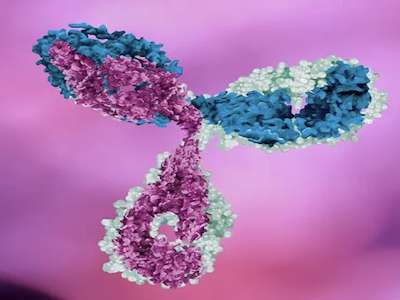

















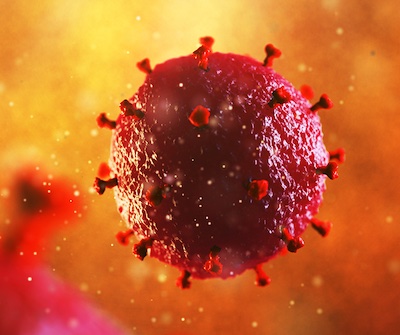





.jpg)
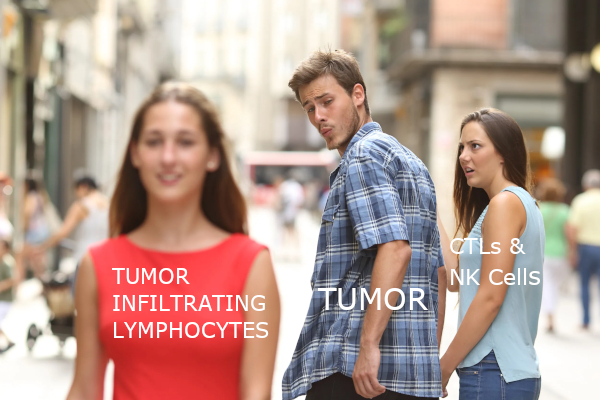









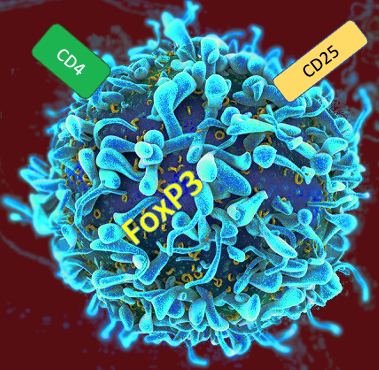
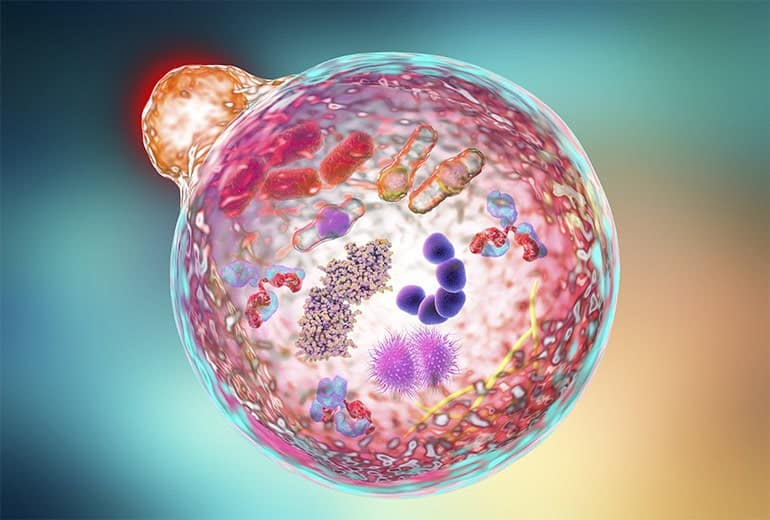
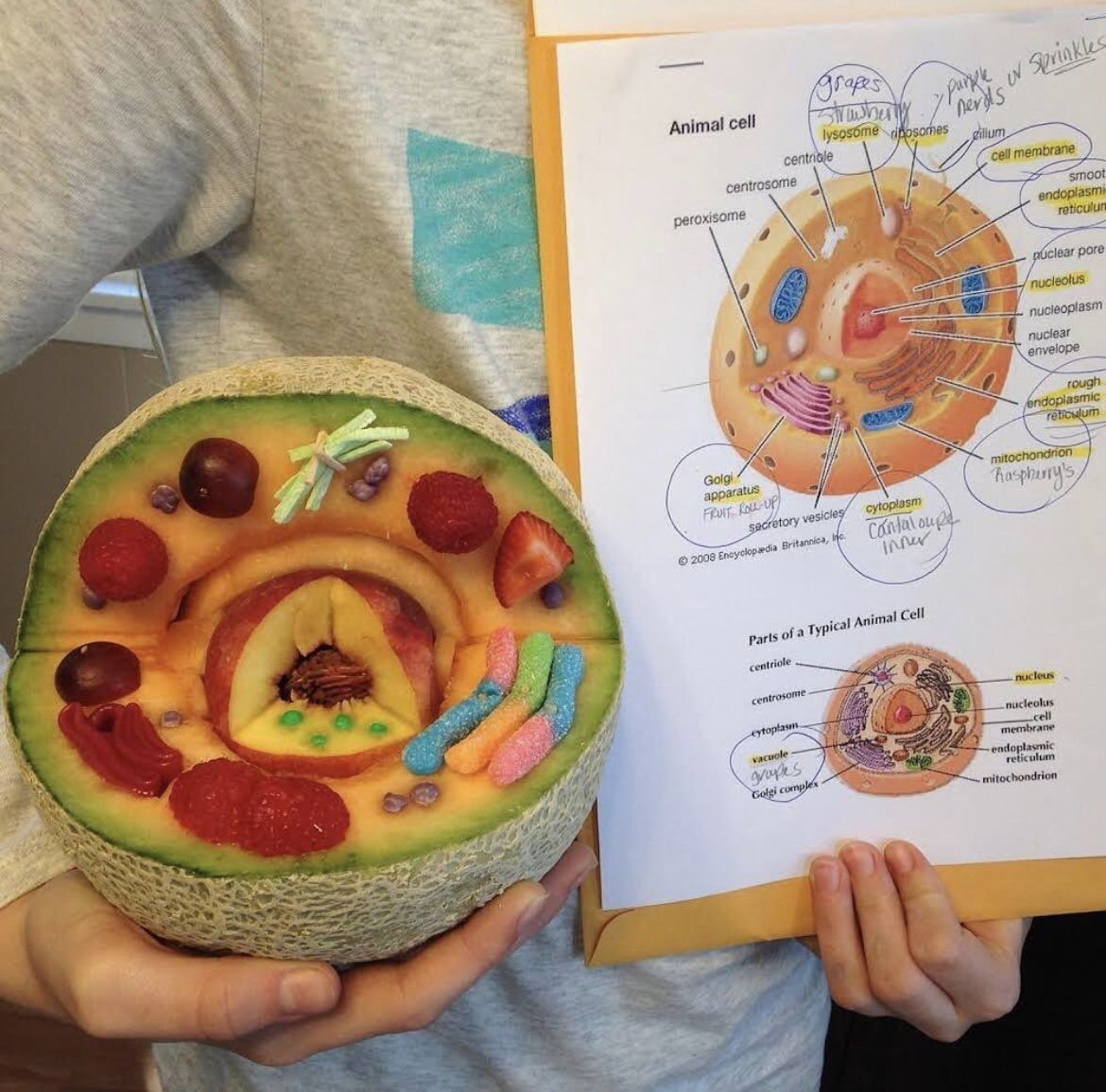






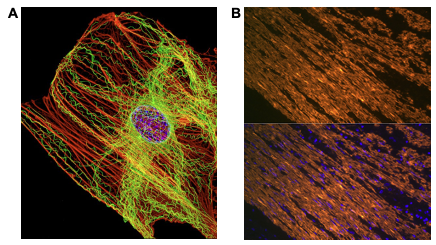
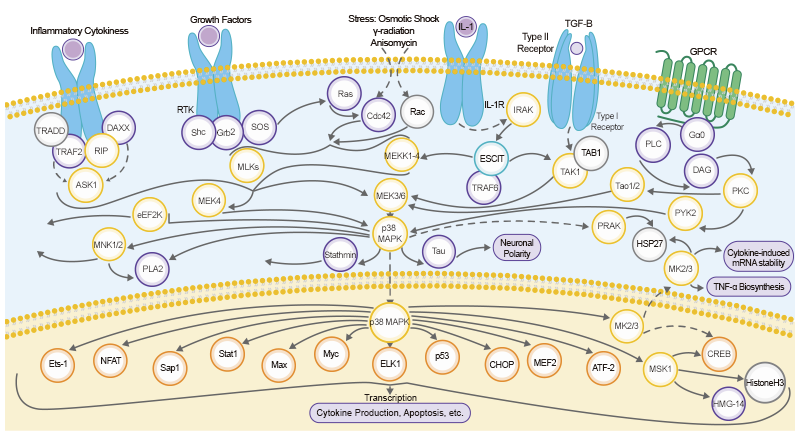


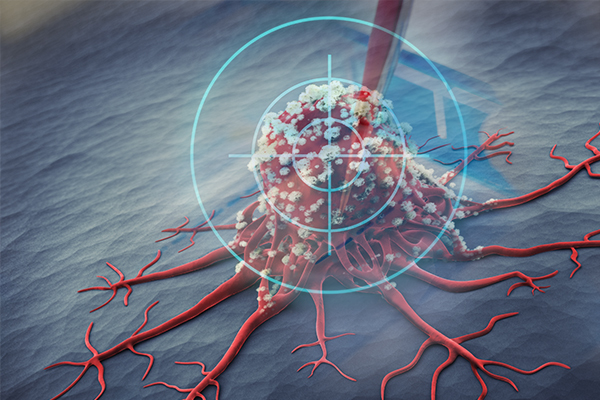
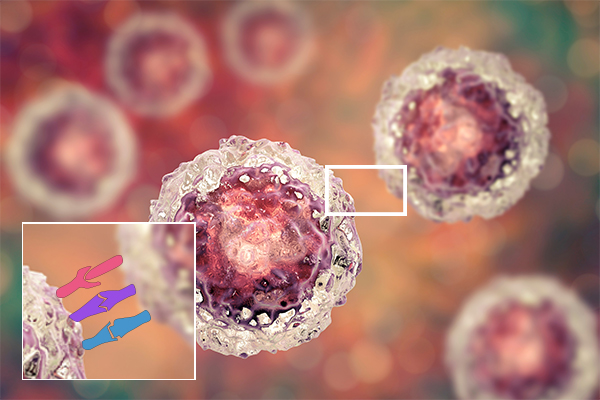
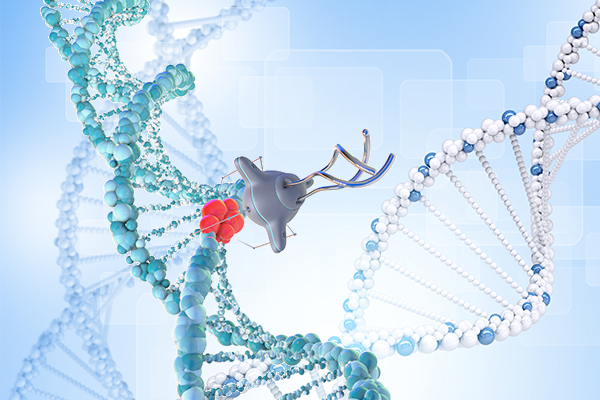
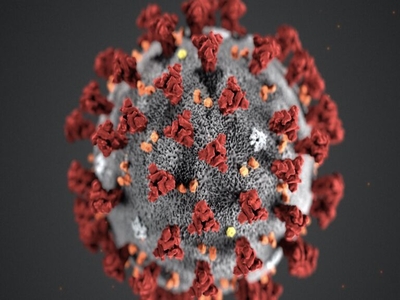

.png)
.png)

.png)
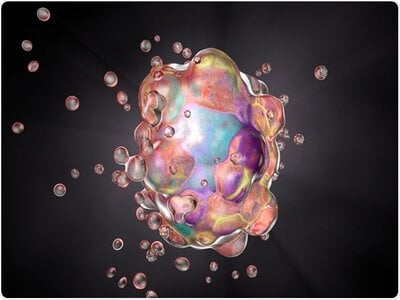



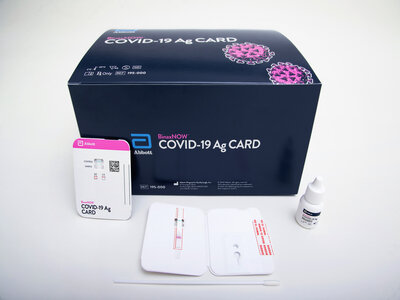









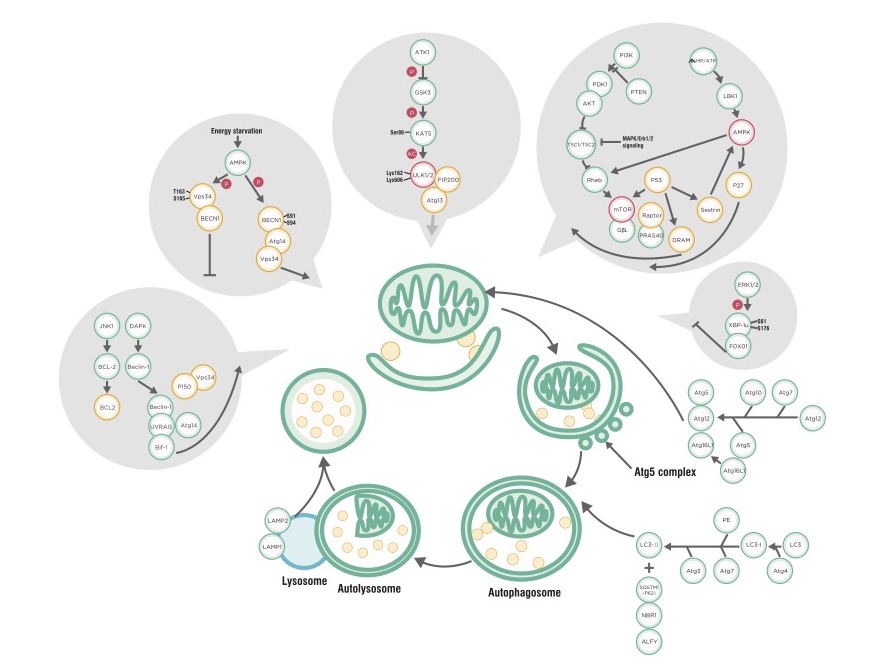
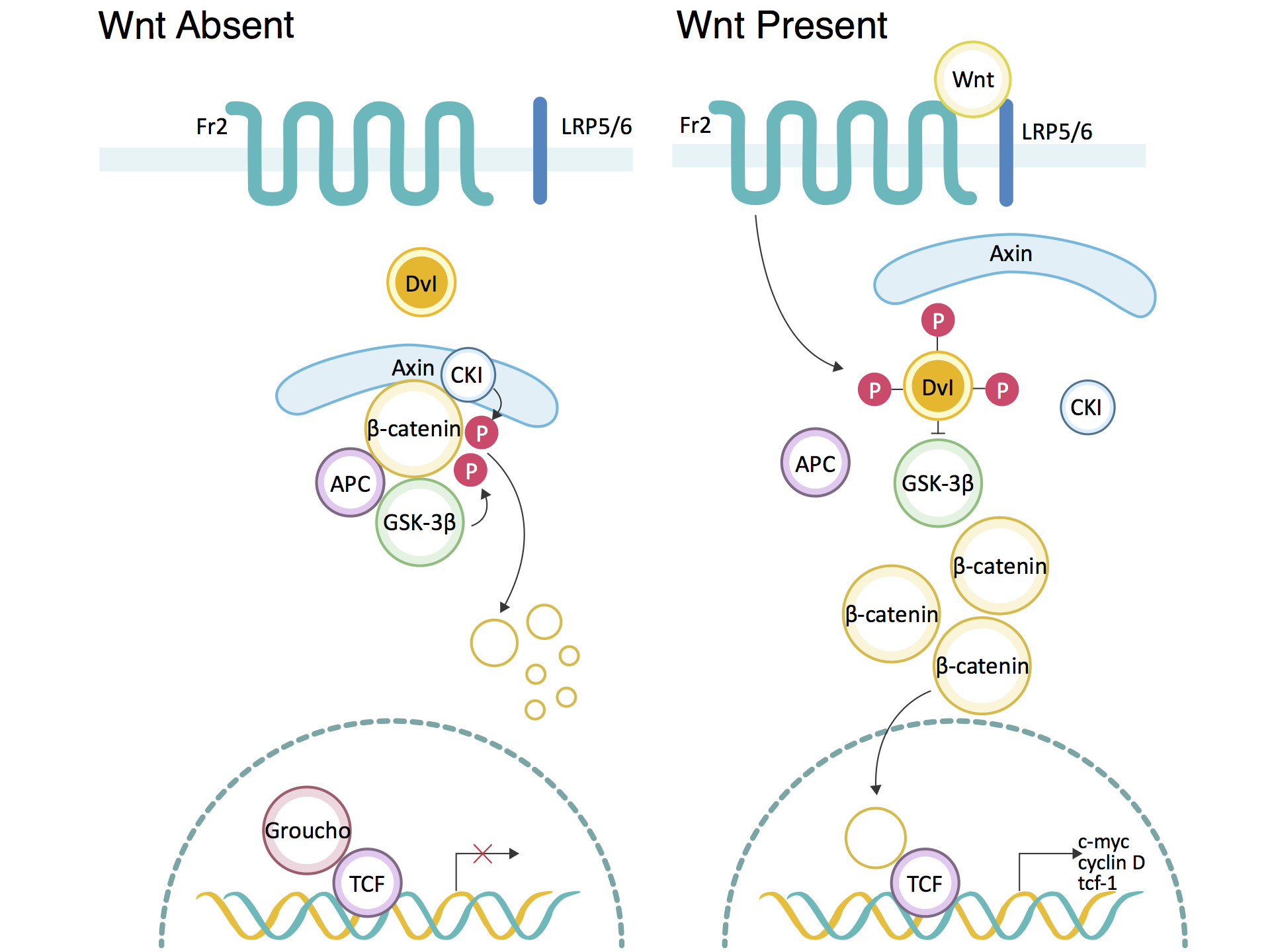





-2.jpg)